Molecular Evolution of Human Adenovirus (HAdV) Species C
- PMID: 30705303
- PMCID: PMC6355881
- DOI: 10.1038/s41598-018-37249-4
Molecular Evolution of Human Adenovirus (HAdV) Species C
Abstract
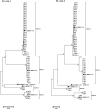
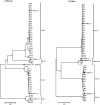
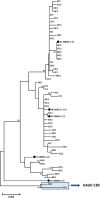
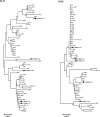
Currently, 88 different Human Adenovirus (HAdV) types are grouped into seven HAdV species A to G. Most types (57) belong to species HAdV-D. Recombination between capsid genes (hexon, penton and fiber) is the main factor contributing to the diversity in species HAdV-D. Noteworthy, species HAdV-C contains so far only five types, although species HAdV-C is highly prevalent and clinically significant in immunosuppressed patients. Therefore, the evolution of species HAdV-C was studied by generating 51 complete genome sequences from circulating strains. Clustering of the whole genome HAdV-C sequences confirmed classical typing results (fifteen HAdV-C1, thirty HAdV-C2, four HAdV-C5, two HAdV-C6). However, two HAdV-C2 strains had a novel penton base sequence and thus were re-labeled as the novel type HAdV-C89. Fiber and early gene region 3 (E3) sequences clustered always with the corresponding prototype sequence but clustering of the E4 region indicated recombination events in 26 out of the 51 sequenced specimens. Recombination of the E1 gene region was detected in 16 circulating strains. As early gene region sequences are not considered in the type definition of HAdVs, evolution of HAdV-C remains on the subtype level. Nonetheless, recombination of the E1 and E4 gene regions may influence the virulence of HAdV-C strains.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

