Invariant phenotype and molecular association of biallelic TET2 mutant myeloid neoplasia
- PMID: 30709865
- PMCID: PMC6373752
- DOI: 10.1182/bloodadvances.2018024216
Invariant phenotype and molecular association of biallelic TET2 mutant myeloid neoplasia
Abstract
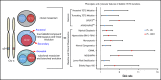
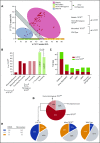
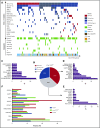
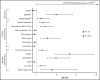
Somatic TET2 mutations (TET2 MT) are frequent in myeloid neoplasia (MN), particularly chronic myelomonocytic leukemia (CMML). TET2 MT includes mostly loss-of-function/hypomorphic hits. Impaired TET2 activity skews differentiation of hematopoietic stem cells toward proliferating myeloid precursors. This study was prompted by the observation of frequent biallelic TET2 gene inactivations (biTET2 i ) in CMML. We speculated that biTET2 i might be associated with distinct clinicohematological features. We analyzed TET2 MT in 1045 patients with MN. Of 82 biTET2 i cases, 66 were biTET2 MT, 13 were hemizygous TET2 MT, and 3 were homozygous TET2 MT (uniparental disomy); the remaining patients (denoted biTET2 - hereafter) were either monoallelic TET2 MT (n = 96) or wild-type TET2 (n = 823). Truncation mutations were found in 83% of biTET2 i vs 65% of biTET2 - cases (P = .02). TET2 hits were founder lesions in 72% of biTET2 i vs 38% of biTET2 - cases (P < .0001). In biTET2 i , significantly concurrent hits included SRSF2 MT (33%; P < .0001) and KRAS/NRAS MT (16%; P = .03) as compared with biTET2 - When the first TET2 hit was ancestral in biTET2 i , the most common subsequent hits affected a second TET2 MT, followed by SRSF2 MT, ASXL1 MT, RAS MT, and DNMT3A MT BiTET2 i patients without any monocytosis showed an absence of SRSF2 MT BiTET2 i patients were older and had monocytosis, CMML, normal karyotypes, and lower-risk disease compared with biTET2 - patients. Hence, while a second TET2 hit occurred frequently, biTET2 i did not portend faster progression but rather determined monocytic differentiation, consistent with its prevalence in CMML. Additionally, biTET2 i showed lower odds of cytopenias and marrow blasts (≥5%) and higher odds of myeloid dysplasia and marrow hypercellularity. Thus, biTET2 i might represent an auxiliary assessment tool in MN.
© 2019 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures

References
-
- Vardiman J, Melo JV, Baccarani M, Thiele J. Chronic myelogenous leukaemia, BCR-ABL1-positive. In: Swerdlow SH, Campo E, Harris NL, et al., eds. WHO Classification of Tumours of Haematopoietic and Lymphoid tissues, 4th ed. Lyon: IARC Press; 2008:32-37.
-
- Yan M, Kanbe E, Peterson LF, et al. . A previously unidentified alternatively spliced isoform of t(8;21) transcript promotes leukemogenesis. Nat Med. 2006;12(8):945-949. - PubMed
-
- Ayton PM, Cleary ML. Molecular mechanisms of leukemogenesis mediated by MLL fusion proteins. Oncogene. 2001;20(40):5695-5707. - PubMed
-
- Fröhling S, Scholl C, Gilliland DG, Levine RL. Genetics of myeloid malignancies: pathogenetic and clinical implications. J Clin Oncol. 2005;23(26):6285-6295. - PubMed
-
- Pabst T, Eyholzer M, Haefliger S, Schardt J, Mueller BU. Somatic CEBPA mutations are a frequent second event in families with germline CEBPA mutations and familial acute myeloid leukemia. J Clin Oncol. 2008;26(31):5088-5093. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

