HLA-C-restricted viral epitopes are associated with an escape mechanism from KIR2DL2+ NK cells in Lassa virus infection
- PMID: 30711514
- PMCID: PMC6413685
- DOI: 10.1016/j.ebiom.2019.01.048
HLA-C-restricted viral epitopes are associated with an escape mechanism from KIR2DL2+ NK cells in Lassa virus infection
Abstract
Background: Lassa virus (LASV) is the etiologic agent of an acute hemorrhagic fever endemic in West Africa. Natural killer (NK) cells control viral infections in part through the interaction between killer cell immunoglobulin-like receptors (KIRs) and their ligands. LASV infection is associated with defective immune responses, including inhibition of NK cell activity in the presence of MHC-class 1+-infected target cells.
Methods: We compared individual KIR and HLA-class 1 genotypes of 68 healthy volunteers to 51 patients infected with LASV in Sierra Leone, including 37 survivors and 14 fatalities. Next, potential HLA-C1, HLA-C2, and HLA-Bw4 binding epitopes were in silico screened among LASV nucleoprotein (NP) and envelope glycoprotein (GP). Selected 10-mer peptides were then tested in peptide-HLA stabilization, KIR binding and polyfunction assays.
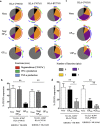
Findings: LASV-infected patients were similar to healthy controls, except for the inhibitory KIR2DL2 gene. We found a specific increase in the HLA-C1:KIR2DL2 interaction in fatalities (10/11) as compared to survivors (12/19) and controls (19/29). We also identified that strong of NP and GP viral epitopes was only observed with HLA-C molecules, and associated with strong inhibition of degranulation in the presence of KIR2DL+ NK cells. This inhibitory effect significantly increased in the presence of the vGP420 variant, detected in 28.1% of LASV sequences.
Interpretation: Our finding suggests that presentation of specific LASV epitopes by HLA-C alleles to the inhibitory KIR2DL2 receptor on NK cells could potentially prevent the killing of infected cells and provides insights into the mechanisms by which LASV can escape NK-cell-mediated immune pressure.
Keywords: HLA-C; KIR-L; Lassa virus; NK cells; Viral escape.
Copyright © 2019. Published by Elsevier B.V.
Figures


References
-
- McCormick J.B., Webb P.A., Krebs J.W., Johnson K.M., Smith E.S. A prospective study of the epidemiology and ecology of Lassa fever. J Inf Dis. 1987;155(3):437–444. - PubMed
-
- Prescott J.B., Marzi A., Safronetz D., Robertson S.J., Feldmann H., Best S.M. Immunobiology of Ebola and Lassa virus infections. Nat Rev Immunol. 2017;17(3):195–207. - PubMed
-
- Ogbu O., Ajuluchukwu E., Uneke C.J. Lassa fever in West African sub-region: an overview. J Vector Borne Dis. 2007;44(1):1–11. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

