Whole-Genome Sequencing for Drug Resistance Profile Prediction in Mycobacterium tuberculosis
- PMID: 30718257
- PMCID: PMC6496161
- DOI: 10.1128/AAC.02175-18
Whole-Genome Sequencing for Drug Resistance Profile Prediction in Mycobacterium tuberculosis
Abstract
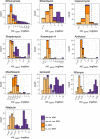
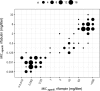
Whole-genome sequencing allows rapid detection of drug-resistant Mycobacterium tuberculosis isolates. However, the availability of high-quality data linking quantitative phenotypic drug susceptibility testing (DST) and genomic data have thus far been limited. We determined drug resistance profiles of 176 genetically diverse clinical M. tuberculosis isolates from the Democratic Republic of the Congo, Ivory Coast, Peru, Thailand, and Switzerland by quantitative phenotypic DST for 11 antituberculous drugs using the BD Bactec MGIT 960 system and 7H10 agar dilution to generate a cross-validated phenotypic DST readout. We compared DST results with predicted drug resistance profiles inferred by whole-genome sequencing. Classification of strains by the two phenotypic DST methods into resistotype/wild-type populations was concordant in 73 to 99% of cases, depending on the drug. Our data suggest that the established critical concentration (5 mg/liter) for ethambutol resistance (MGIT 960 system) is too high and misclassifies strains as susceptible, unlike 7H10 agar dilution. Increased minimal inhibitory concentrations were explained by mutations identified by whole-genome sequencing. Using whole-genome sequences, we were able to predict quantitative drug resistance levels for the majority of drug resistance mutations. Predicting quantitative levels of drug resistance by whole-genome sequencing was partially limited due to incompletely understood drug resistance mechanisms. The overall sensitivity and specificity of whole-genome-based DST were 86.8% and 94.5%, respectively. Despite some limitations, whole-genome sequencing has the potential to infer resistance profiles without the need for time-consuming phenotypic methods.
Keywords: Mycobacterium tuberculosis; drug resistance; drug resistance level prediction; quantitative phenotypic drug susceptibility testing; whole-genome sequencing.
Copyright © 2019 Gygli et al.
Figures


Comment in
-
Whole genome sequencing in Mycobacterium tuberculosis.Ann Transl Med. 2019 Sep;7(Suppl 6):S197. doi: 10.21037/atm.2019.07.28. Ann Transl Med. 2019. PMID: 31656776 Free PMC article. No abstract available.
References
-
- World Health Organization. 2010. Treatment of tuberculosis: guidelines, 4th ed World Health Organization, Geneva, Switzerland.
-
- Domínguez J, Boettger EC, Cirillo D, Cobelens F, Eisenach KD, Gagneux S, Hillemann D, Horsburgh R, Molina-Moya B, Niemann S, Tortoli E, Whitelaw A, Lange C, TBNET, RESIST-TB Networks. 2016. Clinical implications of molecular drug resistance testing for Mycobacterium tuberculosis: a TBNET/RESIST-TB consensus statement. Int J Tuber Lung Dis 20:24–42. doi:10.5588/ijtld.15.0221. - DOI - PubMed
-
- Deggim-Messmer V, Bloemberg GV, Ritter C, Voit A, Hömke R, Keller PM, Böttger EC. 2016. Diagnostic molecular mycobacteriology in regions with low tuberculosis endemicity: combining real-time PCR assays for detection of multiple mycobacterial pathogens with line probe assays for identification of resistance mutations. EBioMedicine 9:228–237. doi:10.1016/j.ebiom.2016.06.016. - DOI - PMC - PubMed
-
- Nathavitharana RR, Hillemann D, Schumacher SG, Schlueter B, Ismail N, Omar SV, Sikhondze W, Havumaki J, Valli E, Boehme C, Denkinger CM. 2016. Multicenter noninferiority evaluation of Hain GenoType MTBDR plus version 2 and Nipro NTM+MDRTB line probe assays for detection of rifampin and isoniazid resistance. J Clin Microbiol 54:1624–1630. doi:10.1128/JCM.00251-16. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

