Genome-wide analyses as part of the international FTLD-TDP whole-genome sequencing consortium reveals novel disease risk factors and increases support for immune dysfunction in FTLD
- PMID: 30739198
- PMCID: PMC6533145
- DOI: 10.1007/s00401-019-01962-9
Genome-wide analyses as part of the international FTLD-TDP whole-genome sequencing consortium reveals novel disease risk factors and increases support for immune dysfunction in FTLD
Abstract
Frontotemporal lobar degeneration with neuronal inclusions of the TAR DNA-binding protein 43 (FTLD-TDP) represents the most common pathological subtype of FTLD. We established the international FTLD-TDP whole-genome sequencing consortium to thoroughly characterize the known genetic causes of FTLD-TDP and identify novel genetic risk factors. Through the study of 1131 unrelated Caucasian patients, we estimated that C9orf72 repeat expansions and GRN loss-of-function mutations account for 25.5% and 13.9% of FTLD-TDP patients, respectively. Mutations in TBK1 (1.5%) and other known FTLD genes (1.4%) were rare, and the disease in 57.7% of FTLD-TDP patients was unexplained by the known FTLD genes. To unravel the contribution of common genetic factors to the FTLD-TDP etiology in these patients, we conducted a two-stage association study comprising the analysis of whole-genome sequencing data from 517 FTLD-TDP patients and 838 controls, followed by targeted genotyping of the most associated genomic loci in 119 additional FTLD-TDP patients and 1653 controls. We identified three genome-wide significant FTLD-TDP risk loci: one new locus at chromosome 7q36 within the DPP6 gene led by rs118113626 (p value = 4.82e - 08, OR = 2.12), and two known loci: UNC13A, led by rs1297319 (p value = 1.27e - 08, OR = 1.50) and HLA-DQA2 led by rs17219281 (p value = 3.22e - 08, OR = 1.98). While HLA represents a locus previously implicated in clinical FTLD and related neurodegenerative disorders, the association signal in our study is independent from previously reported associations. Through inspection of our whole-genome sequence data for genes with an excess of rare loss-of-function variants in FTLD-TDP patients (n ≥ 3) as compared to controls (n = 0), we further discovered a possible role for genes functioning within the TBK1-related immune pathway (e.g., DHX58, TRIM21, IRF7) in the genetic etiology of FTLD-TDP. Together, our study based on the largest cohort of unrelated FTLD-TDP patients assembled to date provides a comprehensive view of the genetic landscape of FTLD-TDP, nominates novel FTLD-TDP risk loci, and strongly implicates the immune pathway in FTLD-TDP pathogenesis.
Keywords: DPP6; HLA; Immunity; TBK1; UNC13A; Whole-genome sequencing FTLD-TDP.
Figures
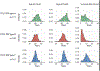

Comment in
-
Genetic risk factors and role of immune dysfunction in FTLD.Nat Rev Neurol. 2019 May;15(5):250-251. doi: 10.1038/s41582-019-0173-5. Nat Rev Neurol. 2019. PMID: 30914791 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
- P30 AG010124/AG/NIA NIH HHS/United States
- R01 DC008552/DC/NIDCD NIH HHS/United States
- U01 AG052943/AG/NIA NIH HHS/United States
- I01 BX003040/BX/BLRD VA/United States
- P30 AG012300/AG/NIA NIH HHS/United States
- RF1 AG051504/AG/NIA NIH HHS/United States
- UH3 NS103870/NS/NINDS NIH HHS/United States
- G0701441/MRC_/Medical Research Council/United Kingdom
- R01 AG041797/AG/NIA NIH HHS/United States
- P30 AG013854/AG/NIA NIH HHS/United States
- R01 NS076837/NS/NINDS NIH HHS/United States
- K01 AG049152/AG/NIA NIH HHS/United States
- K24 AG045333/AG/NIA NIH HHS/United States
- U01 AG058922/AG/NIA NIH HHS/United States
- P50 AG005131/AG/NIA NIH HHS/United States
- P30 AG010133/AG/NIA NIH HHS/United States
- U24 AG021886/AG/NIA NIH HHS/United States
- P50 AG016574/AG/NIA NIH HHS/United States
- P01 AG017586/AG/NIA NIH HHS/United States
- R35 NS097261/NS/NINDS NIH HHS/United States
- P50 AG008702/AG/NIA NIH HHS/United States
- MC_UU_00024/1/MRC_/Medical Research Council/United Kingdom
- U24 NS095871/NS/NINDS NIH HHS/United States
- P01 AG003991/AG/NIA NIH HHS/United States
- P50 AG005681/AG/NIA NIH HHS/United States
- R01 AG051848/AG/NIA NIH HHS/United States
- P01 AG019724/AG/NIA NIH HHS/United States
- R01 AG061796/AG/NIA NIH HHS/United States
- P50 AG025688/AG/NIA NIH HHS/United States
- R01 NS080820/NS/NINDS NIH HHS/United States
- P50 NS072187/NS/NINDS NIH HHS/United States
- MR/J009482/1/MRC_/Medical Research Council/United Kingdom
- P50 AG005133/AG/NIA NIH HHS/United States
- K08 AG052648/AG/NIA NIH HHS/United States
- MR/M023664/1/MRC_/Medical Research Council/United Kingdom
- P01 NS084974/NS/NINDS NIH HHS/United States
- U01 AG046139/AG/NIA NIH HHS/United States
- MR/L016397/1/MRC_/Medical Research Council/United Kingdom
- R01 AG037491/AG/NIA NIH HHS/United States
- U24 NS072026/NS/NINDS NIH HHS/United States
- U01 AG045390/AG/NIA NIH HHS/United States
- P30 AG019610/AG/NIA NIH HHS/United States
- U54 NS092089/NS/NINDS NIH HHS/United States
- R01 NS085770/NS/NINDS NIH HHS/United States
- UG3 NS103870/NS/NINDS NIH HHS/United States
- MR/M008525/1/MRC_/Medical Research Council/United Kingdom
- U01 AG006786/AG/NIA NIH HHS/United States
- P30 NS055077/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

