Feasibility and utility of a panel testing for 114 cancer-associated genes in a clinical setting: A hospital-based study
- PMID: 30742731
- PMCID: PMC6447843
- DOI: 10.1111/cas.13969
Feasibility and utility of a panel testing for 114 cancer-associated genes in a clinical setting: A hospital-based study
Abstract
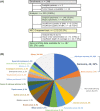
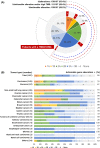
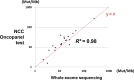
Next-generation sequencing (NGS) of tumor tissue (ie, clinical sequencing) can guide clinical management by providing information about actionable gene aberrations that have diagnostic and therapeutic significance. Here, we undertook a hospital-based prospective study (TOP-GEAR project, 2nd stage) to investigate the feasibility and utility of NGS-based analysis of 114 cancer-associated genes (the NCC Oncopanel test). We examined 230 cases (comprising more than 30 tumor types) of advanced solid tumors, all of which were matched with nontumor samples. Gene profiling data were obtained for 187 cases (81.3%), 111 (59.4%) of which harbored actionable gene aberrations according to the Clinical Practice Guidelines for Next Generation Sequencing in Cancer Diagnosis and Treatment (Edition 1.0) issued by 3 major Japanese cancer-related societies. Twenty-five (13.3%) cases have since received molecular-targeted therapy according to their gene aberrations. These results indicate the utility of tumor-profiling multiplex gene panel testing in a clinical setting in Japan. This study is registered with UMIN Clinical Trials Registry (UMIN 000011141).
Keywords: NCC Oncopanel; actionable gene aberration; clinical sequencing; gene panel test; insurance reimbursement.
© 2019 The Authors. Cancer Science published by John Wiley & Sons Australia, Ltd on behalf of Japanese Cancer Association.
Conflict of interest statement
K.S., T.K., and A.O. are recipients of a collaborative research grant from the Sysmex Corporation. The other authors have no conflicts of interest to declare.
Figures
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

