Investigating the Origins of Membrane Phospholipid Biosynthesis Genes Using Outgroup-Free Rooting
- PMID: 30753429
- PMCID: PMC6431249
- DOI: 10.1093/gbe/evz034
Investigating the Origins of Membrane Phospholipid Biosynthesis Genes Using Outgroup-Free Rooting
Abstract
One of the key differences between Bacteria and Archaea is their canonical membrane phospholipids, which are synthesized by distinct biosynthetic pathways with nonhomologous enzymes. This "lipid divide" has important implications for the early evolution of cells and the type of membrane phospholipids present in the last universal common ancestor. One of the main challenges in studies of membrane evolution is that the key biosynthetic genes are ancient and their evolutionary histories are poorly resolved. This poses major challenges for traditional rooting methods because the only available outgroups are distantly related. Here, we address this issue by using the best available substitution models for single-gene trees, by expanding our analyses to the diversity of uncultivated prokaryotes recently revealed by environmental genomics, and by using two complementary approaches to rooting that do not depend on outgroups. Consistent with some previous analyses, our rooted gene trees support extensive interdomain horizontal transfer of membrane phospholipid biosynthetic genes, primarily from Archaea to Bacteria. They also suggest that the capacity to make archaeal-type membrane phospholipids was already present in last universal common ancestor.
Keywords: lipid divide; lipid evolution; outgroup-free rooting; phylogenetics.
© The Author(s) 2019. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures


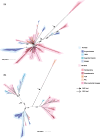
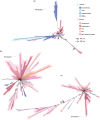
References
-
- Bell SD, Jackson SP.. 1998. Transcription and translation in Archaea: a mosaic of eukaryal and bacterial features. Trends Microbiol. 6(6):222–227. - PubMed
-
- Boucher Y, Kamekura M, Doolittle WF.. 2004. Origins and evolution of isoprenoid lipid biosynthesis in archaea. Mol Microbiol. 52(2):515–527. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

