Expression of STAT3-regulated genes in circulating CD4+ T cells discriminates rheumatoid arthritis independently of clinical parameters in early arthritis
- PMID: 30753680
- PMCID: PMC6587924
- DOI: 10.1093/rheumatology/kez003
Expression of STAT3-regulated genes in circulating CD4+ T cells discriminates rheumatoid arthritis independently of clinical parameters in early arthritis
Abstract
Objectives: Dysregulated signal transduction and activator of transcription-3 (STAT3) signalling in CD4+ T cells has been proposed as an early pathophysiological event in RA. We sought further evidence for this observation, and to determine its clinical relevance.
Methods: Microarray technology was used to measure gene expression in purified peripheral blood CD4+ T cells from treatment-naïve RA patients and disease controls newly recruited from an early arthritis clinic. Analysis focused on 12 previously proposed transcripts, and concurrent STAT3 pathway activation was determined in the same cells by flow cytometry. A pooled analysis of previous and current gene expression findings incorporated detailed clinical parameters and employed multivariate analysis.
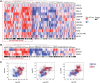
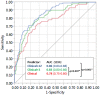
Results: In an independent cohort of 161 patients, expression of 11 of 12 proposed signature genes differed significantly between RA patients and controls, robustly validating the earlier findings. Differential regulation was most pronounced for the STAT3 target genes PIM1, BCL3 and SOCS3 (>1.3-fold difference; P < 0.005), each of whose expression correlated strongly with paired intracellular phospho-STAT3. In a meta-analysis of 279 patients the same three genes accounted for the majority of the signature's ability to discriminate RA patients, which was found to be independent of age, joint involvement or acute phase response.
Conclusion: The STAT3-mediated dysregulation of BCL3, SOCS3 and PIM1 in circulating CD4+ T cells is a discriminatory feature of early RA that occurs independently of acute phase response. The mechanistic and functional implications of this observation at a cellular level warrant clarification.
Keywords: T lymphocytes; gene expression; rheumatoid arthritis.
© The Author(s) 2019. Published by Oxford University Press on behalf of the British Society for Rheumatology.
Figures

References
-
- McInnes IB, O'Dell JR.. State-of-the-art: rheumatoid arthritis. Ann Rheum Dis 2010;69:1898–906. - PubMed
-
- Diogo D, Okada Y, Plenge RM.. Genome-wide association studies to advance our understanding of critical cell types and pathways in rheumatoid arthritis: recent findings and challenges. Curr Opin Rheumatol 2014;26:85–92. - PubMed
-
- Cantaert T, Brouard S, Thurlings RM. et al. Alterations of the synovial T cell repertoire in anti-citrullinated protein antibody-positive rheumatoid arthritis. Arthritis Rheum 2009;60:1944–56. - PubMed
-
- Maxwell LJ, Singh JA.. Abatacept for rheumatoid arthritis: a Cochrane systematic review. J Rheumatol 2010;37:234–45. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

