Delineation of pentatricopeptide repeat codes for target RNA prediction
- PMID: 30753696
- PMCID: PMC6468296
- DOI: 10.1093/nar/gkz075
Delineation of pentatricopeptide repeat codes for target RNA prediction
Erratum in
-
Correction to 'Delineation of pentatricopeptide repeat codes for target RNA prediction'.Nucleic Acids Res. 2025 Jun 20;53(12):gkaf605. doi: 10.1093/nar/gkaf605. Nucleic Acids Res. 2025. PMID: 40548944 Free PMC article. No abstract available.
Abstract
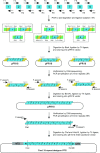
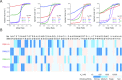
Members of the pentatricopeptide repeat (PPR) protein family are sequence-specific RNA-binding proteins that play crucial roles in organelle RNA metabolism. Each PPR protein consists of a tandem array of PPR motifs, each of which aligns to one nucleotide of the RNA target. The di-residues in the PPR motif, which are referred to as the PPR codes, determine nucleotide specificity. Numerous PPR codes are distributed among the vast number of PPR motifs, but the correlation between PPR codes and RNA bases is poorly understood, which hinders target RNA prediction and functional investigation of PPR proteins. To address this issue, we developed a modular assembly method for high-throughput construction of designer PPRs, and by using this method, 62 designer PPR proteins containing various PPR codes were assembled. Then, the correlation between these PPR codes and RNA bases was systematically explored and delineated. Based on this correlation, the web server PPRCODE (http://yinlab.hzau.edu.cn/pprcode) was developed. Our study will not only serve as a platform for facilitating target RNA prediction and functional investigation of the large number of PPR family proteins but also provide an alternative strategy for the assembly of custom PPRs that can potentially be used for plant organelle RNA manipulation.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
-
Diagnostic test accuracy and cost-effectiveness of tests for codeletion of chromosomal arms 1p and 19q in people with glioma.Cochrane Database Syst Rev. 2022 Mar 2;3(3):CD013387. doi: 10.1002/14651858.CD013387.pub2. Cochrane Database Syst Rev. 2022. PMID: 35233774 Free PMC article.
-
Immunogenicity and seroefficacy of pneumococcal conjugate vaccines: a systematic review and network meta-analysis.Health Technol Assess. 2024 Jul;28(34):1-109. doi: 10.3310/YWHA3079. Health Technol Assess. 2024. PMID: 39046101 Free PMC article.
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
Cited by
-
The Enigmatic Roles of PPR-SMR Proteins in Plants.Adv Sci (Weinh). 2019 May 6;6(13):1900361. doi: 10.1002/advs.201900361. eCollection 2019 Jul 3. Adv Sci (Weinh). 2019. PMID: 31380188 Free PMC article. Review.
-
Pas de Trois: An Overview of Penta-, Tetra-, and Octo-Tricopeptide Repeat Proteins From Chlamydomonas reinhardtii and Their Role in Chloroplast Gene Expression.Front Plant Sci. 2021 Nov 17;12:775366. doi: 10.3389/fpls.2021.775366. eCollection 2021. Front Plant Sci. 2021. PMID: 34868174 Free PMC article. Review.
-
Beyond a PPR-RNA recognition code: Many aspects matter for the multi-targeting properties of RNA editing factor PPR56.PLoS Genet. 2023 Aug 21;19(8):e1010733. doi: 10.1371/journal.pgen.1010733. eCollection 2023 Aug. PLoS Genet. 2023. PMID: 37603555 Free PMC article.
-
Expanding the binding specificity for RNA recognition by a PUF domain.Nat Commun. 2021 Aug 24;12(1):5107. doi: 10.1038/s41467-021-25433-6. Nat Commun. 2021. PMID: 34429425 Free PMC article.
-
GENOMES UNCOUPLED PROTEIN1 binds to plastid RNAs and promotes their maturation.Plant Commun. 2024 Dec 9;5(12):101069. doi: 10.1016/j.xplc.2024.101069. Epub 2024 Aug 22. Plant Commun. 2024. PMID: 39169625 Free PMC article.
References
-
- Barkan A., Small I.. Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 2014; 65:415–442. - PubMed
-
- Wu W., Liu S., Ruwe H., Zhang D., Melonek J., Zhu Y., Hu X., Gusewski S., Yin P., Small I.D. et al.. SOT1, a pentatricopeptide repeat protein with a small MutS-related domain, is required for correct processing of plastid 23S-4.5S rRNA precursors in Arabidopsis thaliana. Plant J. 2016; 85:607–621. - PubMed
-
- Yap A., Kindgren P., Colas des Francs-Small C., Kazama T., Tanz S.K., Toriyama K., Small I.. AEF1/MPR25 is implicated in RNA editing of plastid atpF and mitochondrial nad5, and also promotes atpF splicing in Arabidopsis and rice. Plant J. 2015; 81:661–669. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

