DeepRibo: a neural network for precise gene annotation of prokaryotes by combining ribosome profiling signal and binding site patterns
- PMID: 30753697
- PMCID: PMC6451124
- DOI: 10.1093/nar/gkz061
DeepRibo: a neural network for precise gene annotation of prokaryotes by combining ribosome profiling signal and binding site patterns
Abstract
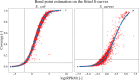
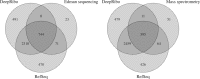
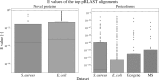
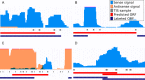
Annotation of gene expression in prokaryotes often finds itself corrected due to small variations of the annotated gene regions observed between different (sub)-species. It has become apparent that traditional sequence alignment algorithms, used for the curation of genomes, are not able to map the full complexity of the genomic landscape. We present DeepRibo, a novel neural network utilizing features extracted from ribosome profiling information and binding site sequence patterns that shows to be a precise tool for the delineation and annotation of expressed genes in prokaryotes. The neural network combines recurrent memory cells and convolutional layers, adapting the information gained from both the high-throughput ribosome profiling data and ribosome binding translation initiation sequence region into one model. DeepRibo is designed as a single model trained on a variety of ribosome profiling experiments, used for the identification of open reading frames in prokaryotes without a priori knowledge of the translational landscape. Through extensive validation of the model trained on various sets of data, multiple species sequence similarity, mass spectrometry and Edman degradation verified proteins, the effectiveness of DeepRibo is highlighted.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
REPARATION: ribosome profiling assisted (re-)annotation of bacterial genomes.Nucleic Acids Res. 2017 Nov 16;45(20):e168. doi: 10.1093/nar/gkx758. Nucleic Acids Res. 2017. PMID: 28977509 Free PMC article.
-
Reannotation of translational start sites in the genome of Mycobacterium tuberculosis.Tuberculosis (Edinb). 2013 Jan;93(1):18-25. doi: 10.1016/j.tube.2012.11.012. Epub 2012 Dec 26. Tuberculosis (Edinb). 2013. PMID: 23273318 Free PMC article.
-
Non-AUG start codons: Expanding and regulating the small and alternative ORFeome.Exp Cell Res. 2020 Jun 1;391(1):111973. doi: 10.1016/j.yexcr.2020.111973. Epub 2020 Mar 21. Exp Cell Res. 2020. PMID: 32209305 Free PMC article. Review.
-
smORFer: a modular algorithm to detect small ORFs in prokaryotes.Nucleic Acids Res. 2021 Sep 7;49(15):e89. doi: 10.1093/nar/gkab477. Nucleic Acids Res. 2021. PMID: 34125903 Free PMC article.
-
The proteome under translational control.Proteomics. 2014 Dec;14(23-24):2647-62. doi: 10.1002/pmic.201400165. Epub 2014 Nov 2. Proteomics. 2014. PMID: 25263132 Review.
Cited by
-
Small proteins in Gram-positive bacteria.FEMS Microbiol Rev. 2023 Nov 1;47(6):fuad064. doi: 10.1093/femsre/fuad064. FEMS Microbiol Rev. 2023. PMID: 38052429 Free PMC article.
-
A Comprehensive Review of Bioinformatics Tools for Genomic Biomarker Discovery Driving Precision Oncology.Genes (Basel). 2024 Aug 6;15(8):1036. doi: 10.3390/genes15081036. Genes (Basel). 2024. PMID: 39202397 Free PMC article. Review.
-
Modelling microbial communities: Harnessing consortia for biotechnological applications.Comput Struct Biotechnol J. 2021 Jul 3;19:3892-3907. doi: 10.1016/j.csbj.2021.06.048. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 34584635 Free PMC article. Review.
-
Lost and Found: Re-searching and Re-scoring Proteomics Data Aids Genome Annotation and Improves Proteome Coverage.mSystems. 2020 Oct 27;5(5):e00833-20. doi: 10.1128/mSystems.00833-20. mSystems. 2020. PMID: 33109751 Free PMC article.
-
Trips-Viz: an environment for the analysis of public and user-generated ribosome profiling data.Nucleic Acids Res. 2021 Jul 2;49(W1):W662-W670. doi: 10.1093/nar/gkab323. Nucleic Acids Res. 2021. PMID: 33950201 Free PMC article.
References
-
- Fields A.P., Rodriguez E.H., Jovanovic M., Stern-Ginossar N., Haas B.J., Mertins P., Raychowdhury R., Hacohen N., Carr S.A., Ingolia N.T. et al. .. A regression-based analysis of ribosome-profiling data reveals a conserved complexity to mammalian translation. Mol. Cell. 2015; 60:816–827. - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

