Cryptococcus deuterogattii VGIIa Infection Associated with Travel to the Pacific Northwest Outbreak Region in an Anti-Granulocyte-Macrophage Colony-Stimulating Factor Autoantibody-Positive Patient in the United States
- PMID: 30755511
- PMCID: PMC6372798
- DOI: 10.1128/mBio.02733-18
Cryptococcus deuterogattii VGIIa Infection Associated with Travel to the Pacific Northwest Outbreak Region in an Anti-Granulocyte-Macrophage Colony-Stimulating Factor Autoantibody-Positive Patient in the United States
Abstract
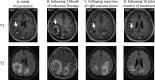
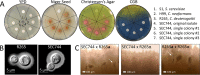
The region encompassing the Pacific Northwest (PNW), Vancouver Island, Oregon, and Washington has been the location of an ongoing Cryptococcus gattii outbreak since the 1990s, and there is evidence that the outbreak is expanding along the West Coast into California. Here we report a clinical case of a 69-year-old, HIV-negative man from North Carolina who was diagnosed with a fungal brain mass by magnetic resonance imaging (MRI) and pathology. He had traveled to Seattle and Vancouver 3 years earlier and to Costa Rica 4 months prior to presentation. Phenotypic evidence showed that the fungal mass isolated from the patient's brain represented C. gattii In agreement with the phenotypic results, multilocus sequence typing (MLST) provided genotypic evidence that assigned the infecting organism within the C. gattii species complex and to the C. deuterogattii VGIIa clade. Whole-genome sequencing revealed >99.99% identity with the C. deuterogattii reference strain R265, indicating that the infecting strain is derived from the highly clonal outbreak strains in the PNW. We conclude that the patient acquired the C. gattii infection during his travel to the region 3 years prior and that the infection was dormant for an extended period of time before causing disease. The patient tested positive for anti-granulocyte-macrophage colony-stimulating factor (GM-CSF) autoantibodies, supporting earlier reports that implicate these autoantibodies as a risk factor associated with C. gattii infection.IMPORTANCE Mortality rates associated with C. gattii infections are estimated to be between 13% and 33%, depending on an individual's predisposition, and C. gattii has caused at least 39 deaths in the PNW region. There have been four other international travel cases reported in patients from Europe and Asia with travel history to the PNW, but this report describes the first North American traveler who acquired C. deuterogattii infection presenting within the United States and the first case of a C. deuterogattii outbreak infection associated with anti-GM-CSF autoantibodies. Early and accurate diagnoses are important for disease prevention and treatment and for control of infectious diseases. Continual reporting of C. deuterogattii infections is necessary to raise awareness of the ongoing outbreak in the PNW and to alert travelers and physicians to the areas of endemicity with potential risks.
Keywords: Cryptococcus deuterogattii; anti-GM-CSF autoantibodies; travel-acquired cryptococcal meningitis.
Copyright © 2019 Applen Clancey et al.
Figures



References
-
- World Health Organization. 1999. Removing obstacles to healthy development. World Health Organization, Geneva, Switzerland.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

