Presence, persistence and effects of pre-treatment HIV-1 drug resistance variants detected using next generation sequencing: A Retrospective longitudinal study from rural coastal Kenya
- PMID: 30759103
- PMCID: PMC6373901
- DOI: 10.1371/journal.pone.0210559
Presence, persistence and effects of pre-treatment HIV-1 drug resistance variants detected using next generation sequencing: A Retrospective longitudinal study from rural coastal Kenya
Abstract
Background: The epidemiology of HIV-1 drug resistance (HIVDR) determined by Sanger capillary sequencing, has been widely studied. However, much less is known about HIVDR detected using next generation sequencing (NGS) methods. We aimed to determine the presence, persistence and effect of pre-treatment HIVDR variants detected using NGS in HIV-1 infected antiretroviral treatment (ART) naïve participants from rural Coastal Kenya.
Methods: In a retrospective longitudinal study, samples from HIV-1 infected participants collected prior [n = 2 time-points] and after [n = 1 time-point] ART initiation were considered. An ultra-deep amplicon-based NGS assay, calling for nucleotide variants at >2.0% frequency of viral population, was used. Suspected virologic failure (sVF) was defined as a one-off HIV-1 viral load of >1000 copies/ml whilst on ART.
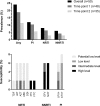
Results: Of the 50 eligible participants, 12 (24.0% [95% CI: 13.1-38.2]) had at least one detectable pre-treatment HIVDR variant against Protease Inhibitors (PIs, n = 6 [12%]), Nucleoside Reverse Transcriptase Inhibitors (NRTIs, n = 4 [8.0%]) and Non-NRTIs (n = 3 [6.0%]). Overall, 15 pre-treatment resistance variants were detected (frequency, range: 2.3-92.0%). A positive correlation was observed between mutation frequency and absolute load for NRTI and/or NNRTI variants (r = 0.761 [p = 0.028]), but not for PI variants (r = -0.117 [p = 0.803]). Participants with pre-treatment NRTI and/or NNRTI resistance had increased odds of sVF (OR = 6.0; 95% CI = 1.0-36.9; p = 0.054).
Conclusions: Using NGS, pre-treatment resistance variants were common, though observed PI variants were unlikely transmitted, but rather probably generated de novo. Even when detected from a low frequency, pre-treatment NRTI and/or NNRTI resistance variants may adversely affect treatment outcomes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Acquired HIV drug resistance among adults living with HIV receiving first-line antiretroviral therapy in Rwanda: A cross-sectional nationally representative survey.Antivir Ther. 2022 Jun;27(3):13596535221102690. doi: 10.1177/13596535221102690. Antivir Ther. 2022. PMID: 35593031 Free PMC article.
-
Minority and majority pretreatment HIV-1 drug resistance associated with failure of first-line nonnucleoside reverse-transcriptase inhibitor antiretroviral therapy in Kenyan women.AIDS. 2019 May 1;33(6):941-951. doi: 10.1097/QAD.0000000000002134. AIDS. 2019. PMID: 30946148 Free PMC article.
-
Study of the impact of HIV genotypic drug resistance testing on therapy efficacy.Verh K Acad Geneeskd Belg. 2001;63(5):447-73. Verh K Acad Geneeskd Belg. 2001. PMID: 11813503 Review.
-
Prevalence and evolution of low frequency HIV drug resistance mutations detected by ultra deep sequencing in patients experiencing first line antiretroviral therapy failure.PLoS One. 2014 Jan 27;9(1):e86771. doi: 10.1371/journal.pone.0086771. eCollection 2014. PLoS One. 2014. PMID: 24475178 Free PMC article.
-
HIV-1 drug resistance and resistance testing.Infect Genet Evol. 2016 Dec;46:292-307. doi: 10.1016/j.meegid.2016.08.031. Epub 2016 Aug 29. Infect Genet Evol. 2016. PMID: 27587334 Free PMC article. Review.
Cited by
-
A cross-sectional study evaluating the frequency of HIV drug resistance mutations among individuals diagnosed with HIV-1 in tenofovir disoproxil fumarate-based pre-exposure prophylaxis rollout programmes in Kenya, Zimbabwe, Eswatini and South Africa.J Int AIDS Soc. 2025 Aug;28(8):e70011. doi: 10.1002/jia2.70011. J Int AIDS Soc. 2025. PMID: 40836601 Free PMC article.
-
High Level of Pre-Treatment HIV-1 Drug Resistance and Its Association with HLA Class I-Mediated Restriction in the Pumwani Sex Worker Cohort.Viruses. 2022 Jan 28;14(2):273. doi: 10.3390/v14020273. Viruses. 2022. PMID: 35215866 Free PMC article.
-
Impact of Low-Frequency Human Immunodeficiency Virus Type 1 Drug Resistance Mutations on Antiretroviral Therapy Outcomes.J Infect Dis. 2024 Jul 25;230(1):86-94. doi: 10.1093/infdis/jiae131. J Infect Dis. 2024. PMID: 39052733 Free PMC article.
-
Low-frequency HIV-1 drug resistance mutations in antiretroviral naïve individuals in Botswana.Medicine (Baltimore). 2022 Jul 15;101(28):e29577. doi: 10.1097/MD.0000000000029577. Medicine (Baltimore). 2022. PMID: 35838991 Free PMC article.
-
Low-Abundance Drug-Resistant HIV-1 Variants in Antiretroviral Drug-Naive Individuals: A Systematic Review of Detection Methods, Prevalence, and Clinical Impact.J Infect Dis. 2020 Apr 27;221(10):1584-1597. doi: 10.1093/infdis/jiz650. J Infect Dis. 2020. PMID: 31809534 Free PMC article.
References
-
- Gupta R.K., et al., Global trends in antiretroviral resistance in treatment-naive individuals with HIV after rollout of antiretroviral treatment in resource-limited settings: a global collaborative study and meta-regression analysis. Lancet, 2012. 380(9849): p. 1250–8. 10.1016/S0140-6736(12)61038-1 - DOI - PMC - PubMed
-
- Hamers R.L., et al., Effect of pretreatment HIV-1 drug resistance on immunological, virological, and drug-resistance outcomes of first-line antiretroviral treatment in sub-Saharan Africa: a multicentre cohort study. Lancet Infect Dis, 2012. 12(4): p. 307–17. 10.1016/S1473-3099(11)70255-9 - DOI - PubMed
-
- Wittkop L., et al., Effect of transmitted drug resistance on virological and immunological response to initial combination antiretroviral therapy for HIV (EuroCoord-CHAIN joint project): a European multicohort study. Lancet Infect Dis, 2011. 11(5): p. 363–71. 10.1016/S1473-3099(11)70032-9 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

