Domestication of Local Microbial Consortia for Efficient Recovery of Gold Through Top-Down Selection in Airlift Bioreactors
- PMID: 30761108
- PMCID: PMC6363673
- DOI: 10.3389/fmicb.2019.00060
Domestication of Local Microbial Consortia for Efficient Recovery of Gold Through Top-Down Selection in Airlift Bioreactors
Abstract
Extreme acidophiles play central roles in the geochemical cycling of diverse elements in low pH environments. This has been harnessed in biotechnologies such as biomining, where microorganisms facilitate the recovery of economically important metals such as gold. By generating both extreme acidity and a chemical oxidant (ferric iron) many species of prokaryotes that thrive in low pH environments not only catalyze mineral dissolution but also trigger both community and individual level adaptive changes. These changes vary in extent and direction depending on the ore mineralogy, water availability and local climate. The use of indigenous versus introduced microbial consortia in biomining practices is still a matter of debate. Yet, indigenous microbial consortia colonizing sulfidic ores that have been domesticated, i.e., selected for their ability to survive under specific polyextreme conditions, are claimed to outperform un-adapted foreign consortia. Despite this, little is known on the domestication of acidic microbial communities and the changes elicited in their members. In this study, high resolution targeted metagenomic techniques were used to analyze the changes occurring in the community structure of local microbial consortia acclimated to growing under extreme acidic conditions and adapted to endure the conditions imposed by the target mineral during biooxidation of a gold concentrate in an airlift reactor over a period of 2 years. The results indicated that operative conditions evolving through biooxidation of the mineral concentrate exerted strong selective pressures that, early on, purge biodiversity in favor of a few Acidithiobacillus spp. over other iron oxidizing acidophiles. Metagenomic analysis of the domesticated consortium present at the end of the adaptation experiment enabled reconstruction of the RVS1-MAG, a novel representative of Acidithiobacillus ferrooxidans from the Andacollo gold mineral district. Comparative genomic analysis performed with this genome draft revealed a net enrichment of gene functions related to heavy metal transport and stress management that are likely to play a significant role in adaptation and survival to adverse conditions experienced by these acidophiles during growth in presence of gold concentrates.
Keywords: Acidithiobacillus; acidophiles; adaptation; consortia; domestication; metagenome derived assembly; targeted metagenomics.
Figures
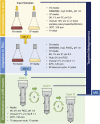
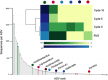
 ); BVS (
); BVS ( ); BVW (
); BVW ( ); SPB (
); SPB ( ); SPS (
); SPS ( ); RVS (
); RVS ( ); PPO (
); PPO ( ); SC: sterile control (-).
); SC: sterile control (-).

Similar articles
-
Convergent Community Assembly among Globally Separated Acidic Cave Biofilms.Appl Environ Microbiol. 2023 Jan 31;89(1):e0157522. doi: 10.1128/aem.01575-22. Epub 2023 Jan 5. Appl Environ Microbiol. 2023. PMID: 36602326 Free PMC article.
-
Microbiological and geochemical dynamics in simulated-heap leaching of a polymetallic sulfide ore.Biotechnol Bioeng. 2008 Nov 1;101(4):739-50. doi: 10.1002/bit.21951. Biotechnol Bioeng. 2008. PMID: 18496880
-
Biooxidation of pyrite by defined mixed cultures of moderately thermophilic acidophiles in pH-controlled bioreactors: significance of microbial interactions.Biotechnol Bioeng. 2004 Sep 5;87(5):574-83. doi: 10.1002/bit.20138. Biotechnol Bioeng. 2004. PMID: 15352055
-
The microbiology of biomining: development and optimization of mineral-oxidizing microbial consortia.Microbiology (Reading). 2007 Feb;153(Pt 2):315-324. doi: 10.1099/mic.0.2006/001206-0. Microbiology (Reading). 2007. PMID: 17259603 Review.
-
Genomic insights into microbial iron oxidation and iron uptake strategies in extremely acidic environments.Environ Microbiol. 2012 Jul;14(7):1597-611. doi: 10.1111/j.1462-2920.2011.02626.x. Epub 2011 Nov 3. Environ Microbiol. 2012. PMID: 22050575 Review.
Cited by
-
Methods for studying microbial acid stress responses: from molecules to populations.FEMS Microbiol Rev. 2024 Sep 18;48(5):fuae015. doi: 10.1093/femsre/fuae015. FEMS Microbiol Rev. 2024. PMID: 38760882 Free PMC article. Review.
-
Recent progress in the application of omics technologies in the study of bio-mining microorganisms from extreme environments.Microb Cell Fact. 2021 Sep 8;20(1):178. doi: 10.1186/s12934-021-01671-7. Microb Cell Fact. 2021. PMID: 34496835 Free PMC article. Review.
-
Influence of mobile genetic elements and insertion sequences in long- and short-term adaptive processes of Acidithiobacillus ferrooxidans strains.Sci Rep. 2023 Jul 5;13(1):10876. doi: 10.1038/s41598-023-37341-4. Sci Rep. 2023. PMID: 37407610 Free PMC article.
-
Integrative Genomics Sheds Light on Evolutionary Forces Shaping the Acidithiobacillia Class Acidophilic Lifestyle.Front Microbiol. 2022 Feb 15;12:822229. doi: 10.3389/fmicb.2021.822229. eCollection 2021. Front Microbiol. 2022. PMID: 35242113 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

