Protein-Protein Affinity Determination by Quantitative FRET Quenching
- PMID: 30765720
- PMCID: PMC6375914
- DOI: 10.1038/s41598-018-35535-9
Protein-Protein Affinity Determination by Quantitative FRET Quenching
Abstract
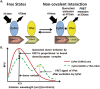
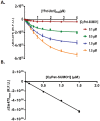
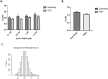
The molecular dissociation constant, Kd, is a well-established parameter to quantitate the affinity of protein-protein or other molecular interactions. Recently, we reported the theoretical basis and experimental procedure for Kd determination using a quantitative FRET method. Here we report a new development of Kd determination by measuring the reduction in donor fluorescence due to acceptor quenching in FRET. A new method of Kd determination was developed from the quantitative measurement of donor fluorescence quenching. The estimated Kd values of SUMO1-Ubc9 interaction based on this method are in good agreement with those determined by other technologies, including FRET acceptor emission. Thus, the acceptor-quenched approach can be used as a complement to the previously developed acceptor excitation method. The new methodology has more general applications regardless whether the acceptor is an excitable fluorophore or a quencher. Thus, these developments provide a complete methodology for protein or other molecule interaction affinity determinations in solution.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Quantitative FRET (qFRET) Technology for the Determination of Protein-Protein Interaction Affinity in Solution.Molecules. 2021 Oct 20;26(21):6339. doi: 10.3390/molecules26216339. Molecules. 2021. PMID: 34770748 Free PMC article. Review.
-
Protein interaction affinity determination by quantitative FRET technology.Biotechnol Bioeng. 2012 Nov;109(11):2875-83. doi: 10.1002/bit.24564. Epub 2012 Jun 18. Biotechnol Bioeng. 2012. PMID: 22711490
-
Development of FRET assay into quantitative and high-throughput screening technology platforms for protein-protein interactions.Ann Biomed Eng. 2011 Apr;39(4):1224-34. doi: 10.1007/s10439-010-0225-x. Epub 2010 Dec 21. Ann Biomed Eng. 2011. PMID: 21174150 Free PMC article.
-
Quantitative FRET (Förster Resonance Energy Transfer) analysis for SENP1 protease kinetics determination.J Vis Exp. 2013 Feb 21;(72):e4430. doi: 10.3791/4430. J Vis Exp. 2013. PMID: 23463095 Free PMC article.
-
A new trend to determine biochemical parameters by quantitative FRET assays.Acta Pharmacol Sin. 2015 Dec;36(12):1408-15. doi: 10.1038/aps.2015.82. Epub 2015 Nov 16. Acta Pharmacol Sin. 2015. PMID: 26567729 Free PMC article. Review.
Cited by
-
Quantitative FRET (qFRET) Technology for the Determination of Protein-Protein Interaction Affinity in Solution.Molecules. 2021 Oct 20;26(21):6339. doi: 10.3390/molecules26216339. Molecules. 2021. PMID: 34770748 Free PMC article. Review.
-
Thermostable homologues of the periplasmic siderophore-binding protein CeuE from Geobacillus stearothermophilus and Parageobacillus thermoglucosidasius.Acta Crystallogr D Struct Biol. 2023 Aug 1;79(Pt 8):694-705. doi: 10.1107/S2059798323004473. Epub 2023 Jul 10. Acta Crystallogr D Struct Biol. 2023. PMID: 37428843 Free PMC article.
-
Mass Spectrometry and Structural Biology Techniques in the Studies on the Coronavirus-Receptor Interaction.Molecules. 2020 Sep 10;25(18):4133. doi: 10.3390/molecules25184133. Molecules. 2020. PMID: 32927621 Free PMC article. Review.
-
Employing 25-Residue Docking Motifs from Modular Polyketide Synthases as Orthogonal Protein Connectors.ACS Synth Biol. 2019 Sep 20;8(9):2017-2024. doi: 10.1021/acssynbio.9b00047. Epub 2019 Sep 12. ACS Synth Biol. 2019. PMID: 31469555 Free PMC article.
-
Human Post-Translational SUMOylation Modification of SARS-CoV-2 Nucleocapsid Protein Enhances Its Interaction Affinity with Itself and Plays a Critical Role in Its Nuclear Translocation.Viruses. 2023 Jul 21;15(7):1600. doi: 10.3390/v15071600. Viruses. 2023. PMID: 37515286 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

