Error-Prone Replication through UV Lesions by DNA Polymerase θ Protects against Skin Cancers
- PMID: 30773314
- PMCID: PMC6453116
- DOI: 10.1016/j.cell.2019.01.023
Error-Prone Replication through UV Lesions by DNA Polymerase θ Protects against Skin Cancers
Abstract
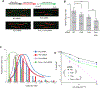
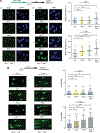
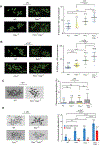
Cancers from sun-exposed skin accumulate "driver" mutations, causally implicated in oncogenesis. Because errors incorporated during translesion synthesis (TLS) opposite UV lesions would generate these mutations, TLS mechanisms are presumed to underlie cancer development. To address the role of TLS in skin cancer formation, we determined which DNA polymerase is responsible for generating UV mutations, analyzed the relative contributions of error-free TLS by Polη and error-prone TLS by Polθ to the replication of UV-damaged DNA and to genome stability, and examined the incidence of UV-induced skin cancers in Polθ-/-, Polη-/-, and Polθ-/- Polη-/- mice. Our findings that the incidence of skin cancers rises in Polθ-/- mice and is further exacerbated in Polθ-/- Polη-/- mice compared with Polη-/- mice support the conclusion that error-prone TLS by Polθ provides a safeguard against tumorigenesis and suggest that cancer formation can ensue in the absence of somatic point mutations.
Keywords: DNA polymerase η; DNA polymerase θ; UV lesions; UV signature mutations; error-free translesion synthesis; error-prone translesion synthesis; genomic rearrangements; replication stress; replication through UV lesions; skin cancers.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing interests.
Figures


Comment in
-
An Error-Prone Polymerase in the Fight against Cancer.Cell. 2019 Mar 7;176(6):1241-1243. doi: 10.1016/j.cell.2019.02.011. Cell. 2019. PMID: 30849368
Similar articles
-
Requirement for functional DNA polymerase eta in genome-wide repair of UV-induced DNA damage during S phase.DNA Repair (Amst). 2010 Jul 1;9(7):754-64. doi: 10.1016/j.dnarep.2010.03.013. Epub 2010 Apr 24. DNA Repair (Amst). 2010. PMID: 20457011
-
UV-induced mutations in epidermal cells of mice defective in DNA polymerase η and/or ι.DNA Repair (Amst). 2015 May;29:139-46. doi: 10.1016/j.dnarep.2015.02.006. Epub 2015 Feb 16. DNA Repair (Amst). 2015. PMID: 25733082
-
Interaction with DNA polymerase eta is required for nuclear accumulation of REV1 and suppression of spontaneous mutations in human cells.DNA Repair (Amst). 2009 May 1;8(5):585-99. doi: 10.1016/j.dnarep.2008.12.006. Epub 2009 Jan 21. DNA Repair (Amst). 2009. PMID: 19157994
-
Xeroderma pigmentosum variant and error-prone DNA polymerases.Biochimie. 2003 Nov;85(11):1123-32. doi: 10.1016/j.biochi.2003.10.009. Biochimie. 2003. PMID: 14726018 Review.
-
Filling gaps in translesion DNA synthesis in human cells.Mutat Res Genet Toxicol Environ Mutagen. 2018 Dec;836(Pt B):127-142. doi: 10.1016/j.mrgentox.2018.02.004. Epub 2018 Feb 23. Mutat Res Genet Toxicol Environ Mutagen. 2018. PMID: 30442338 Review.
Cited by
-
Discovery of a small-molecule inhibitor that traps Polθ on DNA and synergizes with PARP inhibitors.Nat Commun. 2024 Apr 5;15(1):2862. doi: 10.1038/s41467-024-46593-1. Nat Commun. 2024. PMID: 38580648 Free PMC article.
-
Canonical and Non-Canonical Roles of Human DNA Polymerase η.Genes (Basel). 2024 Sep 27;15(10):1271. doi: 10.3390/genes15101271. Genes (Basel). 2024. PMID: 39457395 Free PMC article. Review.
-
Translesion synthesis DNA polymerases η, ι, and ν promote mutagenic replication through the anticancer nucleoside cytarabine.J Biol Chem. 2019 Dec 13;294(50):19048-19054. doi: 10.1074/jbc.RA119.011381. Epub 2019 Nov 4. J Biol Chem. 2019. PMID: 31685662 Free PMC article.
-
Mechanisms of UV-induced mutations and skin cancer.Genome Instab Dis. 2020 May;1(3):99-113. doi: 10.1007/s42764-020-00009-8. Epub 2020 Mar 19. Genome Instab Dis. 2020. PMID: 34589668 Free PMC article.
-
Structural basis of error-prone DNA synthesis by DNA polymerase θ.Nat Commun. 2025 Feb 28;16(1):2063. doi: 10.1038/s41467-025-57269-9. Nat Commun. 2025. PMID: 40021647 Free PMC article.
References
-
- Besaratinia A, and Pfeifer GP (2006). Investigating human cancer etiology by DNA lesion footprinting and mutagenicity analysis. Carcinogenesis 27, 1526–1537. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

