Determining 3'-Termini and Sequences of Nascent Single-Stranded Viral DNA Molecules during HIV-1 Reverse Transcription in Infected Cells
- PMID: 30774124
- PMCID: PMC6682491
- DOI: 10.3791/58715
Determining 3'-Termini and Sequences of Nascent Single-Stranded Viral DNA Molecules during HIV-1 Reverse Transcription in Infected Cells
Abstract
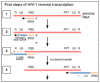
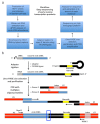
Monitoring of nucleic acid intermediates during virus replication provides insights into the effects and mechanisms of action of antiviral compounds and host cell proteins on viral DNA synthesis. Here we address the lack of a cell-based, high-coverage, and high-resolution assay that is capable of defining retroviral reverse transcription intermediates within the physiological context of virus infection. The described method captures the 3'-termini of nascent complementary DNA (cDNA) molecules within HIV-1 infected cells at single nucleotide resolution. The protocol involves harvesting of whole cell DNA, targeted enrichment of viral DNA via hybrid capture, adaptor ligation, size fractionation by gel purification, PCR amplification, deep sequencing, and data analysis. A key step is the efficient and unbiased ligation of adaptor molecules to open 3'-DNA termini. Application of the described method determines the abundance of reverse transcripts of each particular length in a given sample. It also provides information about the (internal) sequence variation in reverse transcripts and thereby any potential mutations. In general, the assay is suitable for any questions relating to DNA 3'-extension, provided that the template sequence is known.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
