Identification of functional long non-coding RNAs in C. elegans
- PMID: 30777050
- PMCID: PMC6378714
- DOI: 10.1186/s12915-019-0635-7
Identification of functional long non-coding RNAs in C. elegans
Abstract
Background: Functional characterisation of the compact genome of the model organism Caenorhabditis elegans remains incomplete despite its sequencing 20 years ago. The last decade of research has seen a tremendous increase in the number of non-coding RNAs identified in various organisms. While we have mechanistic understandings of small non-coding RNA pathways, long non-coding RNAs represent a diverse class of active transcripts whose function remains less well characterised.
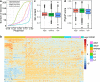
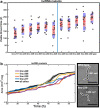
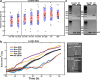
Results: By analysing hundreds of published transcriptome datasets, we annotated 3392 potential lncRNAs including 143 multi-exonic loci that showed increased nucleotide conservation and GC content relative to other non-coding regions. Using CRISPR/Cas9 genome editing, we generated deletion mutants for ten long non-coding RNA loci. Using automated microscopy for in-depth phenotyping, we show that six of the long non-coding RNA loci are required for normal development and fertility. Using RNA interference-mediated gene knock-down, we provide evidence that for two of the long non-coding RNA loci, the observed phenotypes are dependent on the corresponding RNA transcripts.
Conclusions: Our results highlight that a large section of the non-coding regions of the C. elegans genome remains unexplored. Based on our in vivo analysis of a selection of high-confidence lncRNA loci, we expect that a significant proportion of these high-confidence regions is likely to have a biological function at either the genomic or the transcript level.
Keywords: C. elegans; CRISPR; Long non-coding RNA; Non-coding; lincRNA; lncRNA.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare that they have no competing interests. The datasets supporting the conclusions of this article are available in the NCBI SRA repository (
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
- MC_UU_00007/15/MRC_/Medical Research Council/United Kingdom
- 104640/Z/14/Z/WT_/Wellcome Trust/United Kingdom
- 092096/Z/10/Z/WT_/Wellcome Trust/United Kingdom
- MC_UU_12008/1/MRC_/Medical Research Council/United Kingdom
- 18583/CRUK_/Cancer Research UK/United Kingdom
- BBS/E/T/000PR9783/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BBS/E/T/000PR9817/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- C13474/A18583/CRUK_/Cancer Research UK/United Kingdom
- WT_/Wellcome Trust/United Kingdom
- 104640/Z/14/Z/WT_/Wellcome Trust/United Kingdom
- C6946/A14492/CRUK_/Cancer Research UK/United Kingdom
- BBS/E/T/000PR9818/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MR/P026028/1/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Miscellaneous

