Eliciting the impacts of cellular noise on metabolic trade-offs by quantitative mass imaging
- PMID: 30783105
- PMCID: PMC6381102
- DOI: 10.1038/s41467-019-08717-w
Eliciting the impacts of cellular noise on metabolic trade-offs by quantitative mass imaging
Abstract
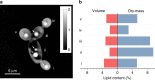
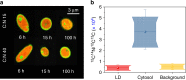
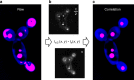
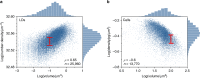
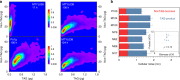
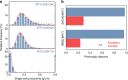
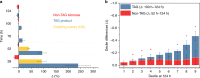
Optimal metabolic trade-offs between growth and productivity are key constraints in strain optimization by metabolic engineering; however, how cellular noise impacts these trade-offs and drives the emergence of subpopulations with distinct resource allocation strategies, remains largely unknown. Here, we introduce a single-cell strategy for quantifying the trade-offs between triacylglycerol production and growth in the oleaginous microorganism Yarrowia lipolytica. The strategy relies on high-throughput quantitative-phase imaging and, enabled by nanoscale secondary ion mass spectrometry analyses and dedicated image processing, allows us to image how resources are partitioned between growth and productivity. Enhanced precision over population-averaging biotechnologies and conventional microscopy demonstrates how cellular noise impacts growth and productivity differently. As such, subpopulations with distinct metabolic trade-offs emerge, with notable impacts on strain performance and robustness. By quantifying the self-degradation of cytosolic macromolecules under nutrient-limiting conditions, we discover the cell-to-cell heterogeneity in protein and fatty-acid recycling, unmasking a potential bet-hedging strategy under starvation.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Characterization of a lipid droplet protein from Yarrowia lipolytica that is required for its oleaginous phenotype.Biochim Biophys Acta Mol Cell Biol Lipids. 2018 Oct;1863(10):1193-1205. doi: 10.1016/j.bbalip.2018.07.010. Epub 2018 Jul 25. Biochim Biophys Acta Mol Cell Biol Lipids. 2018. PMID: 30053597
-
D-stat culture for studying the metabolic shifts from oxidative metabolism to lipid accumulation and citric acid production in Yarrowia lipolytica.J Biotechnol. 2014 Jan 20;170:35-41. doi: 10.1016/j.jbiotec.2013.11.008. Epub 2013 Dec 4. J Biotechnol. 2014. PMID: 24316225
-
Laboratory evolution strategies for improving lipid accumulation in Yarrowia lipolytica.Appl Microbiol Biotechnol. 2019 Oct;103(20):8585-8596. doi: 10.1007/s00253-019-10088-7. Epub 2019 Sep 11. Appl Microbiol Biotechnol. 2019. PMID: 31511932
-
Holistic Approaches in Lipid Production by Yarrowia lipolytica.Trends Biotechnol. 2018 Nov;36(11):1157-1170. doi: 10.1016/j.tibtech.2018.06.007. Epub 2018 Jul 11. Trends Biotechnol. 2018. PMID: 30006239 Review.
-
Yarrowia lipolytica: A model and a tool to understand the mechanisms implicated in lipid accumulation.Biochimie. 2009 Jun;91(6):692-6. doi: 10.1016/j.biochi.2009.02.004. Epub 2009 Feb 25. Biochimie. 2009. PMID: 19248816 Review.
Cited by
-
Deep learning classification of lipid droplets in quantitative phase images.PLoS One. 2021 Apr 5;16(4):e0249196. doi: 10.1371/journal.pone.0249196. eCollection 2021. PLoS One. 2021. PMID: 33819277 Free PMC article.
-
Integrative quantitative-phase and airy light-sheet imaging.Sci Rep. 2020 Nov 19;10(1):20150. doi: 10.1038/s41598-020-76730-x. Sci Rep. 2020. PMID: 33214600 Free PMC article.
-
Microbial metabolic noise.WIREs Mech Dis. 2021 May;13(3):e1512. doi: 10.1002/wsbm.1512. Epub 2020 Nov 23. WIREs Mech Dis. 2021. PMID: 33225608 Free PMC article. Review.
-
Noise suppression in stochastic genetic circuits using PID controllers.PLoS Comput Biol. 2021 Jul 28;17(7):e1009249. doi: 10.1371/journal.pcbi.1009249. eCollection 2021 Jul. PLoS Comput Biol. 2021. PMID: 34319990 Free PMC article.
-
Synthetic aperture interference light (SAIL) microscopy for high-throughput label-free imaging.Appl Phys Lett. 2021 Dec 6;119(23):233701. doi: 10.1063/5.0065628. Epub 2021 Dec 8. Appl Phys Lett. 2021. PMID: 34924588 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

