Cytosine-Based TET Enzyme Inhibitors
- PMID: 30783500
- PMCID: PMC6378777
- DOI: 10.1021/acsmedchemlett.8b00474
Cytosine-Based TET Enzyme Inhibitors
Abstract
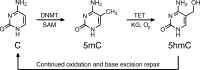
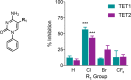
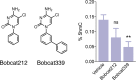
DNA methylation is known as the prima donna epigenetic mark for its critical role in regulating local gene transcription. Changes in the landscape of DNA methylation across the genome occur during cellular transition, such as differentiation and altered neuronal plasticity, and become dysregulated in disease states such as cancer. The TET family of enzymes is known to be responsible for catalyzing the reverse process that is DNA demethylation by recognizing 5-methylcytosine and oxidizing the methyl group via an Fe(II)/alpha-ketoglutarate-dependent mechanism. Here, we describe the design, synthesis, and evaluation of novel cytosine-based TET enzyme inhibitors, a class of small molecule probes previously underdeveloped but broadly desired in the field of epigenetics. We identify a promising cytosine-based lead compound, Bobcat339, that has mid-μM inhibitor activity against TET1 and TET2, but does not inhibit the DNA methyltransferase, DNMT3a. In silico modeling of the TET enzyme active site is used to rationalize the activity of Bobcat339 and other cytosine-based inhibitors. These new molecular tools will be useful to the field of epigenetics and serve as a starting point for new therapeutics that target DNA methylation and gene transcription.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

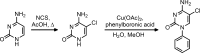

Similar articles
-
Small Molecule Inhibitors of TET Dioxygenases: Bobcat339 Activity Is Mediated by Contaminating Copper(II).ACS Med Chem Lett. 2022 Apr 21;13(5):792-798. doi: 10.1021/acsmedchemlett.1c00677. eCollection 2022 May 12. ACS Med Chem Lett. 2022. PMID: 35586434 Free PMC article.
-
TET enzymes and DNA hydroxymethylation in neural development and function - how critical are they?Genomics. 2014 Nov;104(5):334-40. doi: 10.1016/j.ygeno.2014.08.018. Epub 2014 Sep 6. Genomics. 2014. PMID: 25200796 Review.
-
Tet Enzymes, Variants, and Differential Effects on Function.Front Cell Dev Biol. 2018 Mar 5;6:22. doi: 10.3389/fcell.2018.00022. eCollection 2018. Front Cell Dev Biol. 2018. PMID: 29556496 Free PMC article. Review.
-
Stable Oxidative Cytosine Modifications Accumulate in Cardiac Mesenchymal Cells From Type2 Diabetes Patients: Rescue by α-Ketoglutarate and TET-TDG Functional Reactivation.Circ Res. 2018 Jan 5;122(1):31-46. doi: 10.1161/CIRCRESAHA.117.311300. Epub 2017 Nov 20. Circ Res. 2018. PMID: 29158345
-
How Human TET2 Enzyme Catalyzes the Oxidation of Unnatural Cytosine Modifications in Double-Stranded DNA.ACS Catal. 2022 May 6;12(9):5327-5344. doi: 10.1021/acscatal.2c00024. Epub 2022 Apr 19. ACS Catal. 2022. PMID: 36339349 Free PMC article.
Cited by
-
SAR insights into TET2 catalytic domain inhibition: Synthesis of 2-Hydroxy-4-Methylene-pentanedicarboxylates.Bioorg Med Chem. 2021 Jun 1;39:116141. doi: 10.1016/j.bmc.2021.116141. Epub 2021 Apr 20. Bioorg Med Chem. 2021. PMID: 33894507 Free PMC article.
-
Small Molecule Inhibitors of TET Dioxygenases: Bobcat339 Activity Is Mediated by Contaminating Copper(II).ACS Med Chem Lett. 2022 Apr 21;13(5):792-798. doi: 10.1021/acsmedchemlett.1c00677. eCollection 2022 May 12. ACS Med Chem Lett. 2022. PMID: 35586434 Free PMC article.
-
Epigenetic Regulators of DNA Cytosine Modification: Promising Targets for Cancer Therapy.Biomedicines. 2023 Feb 21;11(3):654. doi: 10.3390/biomedicines11030654. Biomedicines. 2023. PMID: 36979633 Free PMC article. Review.
-
E2F1 and epigenetic modifiers orchestrate breast cancer progression by regulating oxygen-dependent ESRP1 expression.Oncogenesis. 2021 Aug 6;10(8):58. doi: 10.1038/s41389-021-00347-6. Oncogenesis. 2021. PMID: 34362878 Free PMC article.
-
TET activity safeguards pluripotency throughout embryonic dormancy.Nat Struct Mol Biol. 2024 Oct;31(10):1625-1639. doi: 10.1038/s41594-024-01313-7. Epub 2024 May 23. Nat Struct Mol Biol. 2024. PMID: 38783076 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

