Offspring Microbiomes Differ Across Breeding Sites in a Panmictic Species
- PMID: 30787910
- PMCID: PMC6372503
- DOI: 10.3389/fmicb.2019.00035
Offspring Microbiomes Differ Across Breeding Sites in a Panmictic Species
Abstract
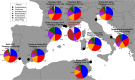
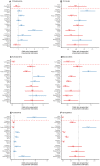
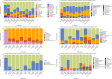
High dispersal rates are known to homogenize host's population genetic structure in panmictic species and to disrupt host local adaptation to the environment. Long-distance dispersal might also spread micro-organisms across large geographical areas. However, so far, to which extent selection mechanisms that shape host's population genetics are mirrored in the population structure of the enteric microbiome remains unclear. High dispersal rates and horizontal parental transfer may homogenize bacterial communities between breeding sites (homogeneous hypothesis). Alternatively, strong selection from the local environment may differentiate bacterial communities between breeding sites (heterogeneous hypothesis). Furthermore, selection from age-specific environmental or physiological factors may differentiate the microbiome between juveniles and adults. Here, we analyzed the cloacal bacterial 16S rRNA gene of fledgling greater flamingos, Phoenicopterus roseus, across nine western Mediterranean breeding sites and four breeding seasons (n = 731) and adult birds (n = 27) from a single site. We found that fledgling cloacal microbiome, as measured by alpha diversity, beta diversity, the relative abundance of assigned sequence variants (ASVs) belonging to a phylum and genus composition within phylum, varied significantly between sampling sites and across time within site despite high adult dispersal rates. The spatio-temporal effects were stronger on individual ASV absence/presence than on ASV abundance (i.e., than on core microbiome composition). Spatial effects had a stronger effect than temporal effects, particularly on ASV abundance. Our study supports the heterogeneous hypothesis whereby local environmental conditions select and differentiate bacterial communities, thus countering the homogenizing effects of high-dispersing host species. In addition, differences in core microbiome between adult vs. fledgling samples suggests that differences in age-specific environmental and/or physiological factors result in differential selection pressure of core enteric microbiome between age classes, even within the same environment. In particular, the genus Corynebacterium, associated with both seasonal fat uptake and migration in previous studies, was much more abundant in high-dispersing fledglings than in more resident adults. To conclude, selection mechanisms that shape the host's genetic structure cannot be extended to the genetic structure of the enteric microbiome, which has important implications regarding our understanding of both host local adaptation mechanisms and enteric microbiome population genetics.
Keywords: Phoenicopterus roseus; dispersal; greater flamingos; gut microbiome; population differentiation.
Figures


References
-
- Altizer S., Harvell D., Friedle E. (2003). Rapid evolutionary dynamics and disease threats to biodiversity. Trends Ecol. Evol. 18 589–596. 10.1016/j.tree.2003.08.013 - DOI
-
- Amat J. A., Hortas F., Arroyo G. M., Rendón M. A., Ramírez J. M., Rendón-Martos M., et al. (2007). Interannual variations in feeding frequencies and food quality of greater flamingo chicks (Phoenicopterus roseus): evidence from plasma chemistry and effects on body condition. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 147 569–576. 10.1016/j.cbpa.2007.02.006 - DOI - PubMed
-
- Amat J. A., Rendón M. A., Rendón-Martos M., Garrido A., Ramírez J. M. (2005). Ranging behaviour of greater flamingos during the breeding and post-breeding periods: Linking connectivity to biological processes. Biol. Conserv. 125 183–192. 10.1016/j.biocon.2005.02.018 - DOI
LinkOut - more resources
Full Text Sources

