Human colon organoids reveal distinct physiologic and oncogenic Wnt responses
- PMID: 30792186
- PMCID: PMC6400532
- DOI: 10.1084/jem.20180823
Human colon organoids reveal distinct physiologic and oncogenic Wnt responses
Abstract
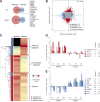
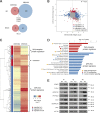
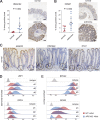
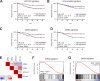
Constitutive Wnt activation upon loss of Adenoma polyposis coli (APC) acts as main driver of colorectal cancer (CRC). Targeting Wnt signaling has proven difficult because the pathway is crucial for homeostasis and stem cell renewal. To distinguish oncogenic from physiological Wnt activity, we have performed transcriptome and proteome profiling in isogenic human colon organoids. Culture in the presence or absence of exogenous ligand allowed us to discriminate receptor-mediated signaling from the effects of CRISPR/Cas9-induced APC loss. We could catalog two nonoverlapping molecular signatures that were stable at distinct levels of stimulation. Newly identified markers for normal stem/progenitor cells and adenomas were validated by immunohistochemistry and flow cytometry. We found that oncogenic Wnt signals are associated with good prognosis in tumors of the consensus molecular subtype 2 (CMS2). In contrast, receptor-mediated signaling was linked to CMS4 tumors and poor prognosis. Together, our data represent a valuable resource for biomarkers that allow more precise stratification of Wnt responses in CRC.
© 2019 Michels et al.
Figures





Similar articles
-
Wnt-driven LARGE2 mediates laminin-adhesive O-glycosylation in human colonic epithelial cells and colorectal cancer.Cell Commun Signal. 2020 Jun 25;18(1):102. doi: 10.1186/s12964-020-00561-6. Cell Commun Signal. 2020. PMID: 32586342 Free PMC article.
-
In Colorectal Cancer Cells With Mutant KRAS, SLC25A22-Mediated Glutaminolysis Reduces DNA Demethylation to Increase WNT Signaling, Stemness, and Drug Resistance.Gastroenterology. 2020 Dec;159(6):2163-2180.e6. doi: 10.1053/j.gastro.2020.08.016. Epub 2020 Aug 16. Gastroenterology. 2020. PMID: 32814111
-
Wnt signaling and colon tumorigenesis--a view from the periphery.Exp Cell Res. 2011 Nov 15;317(19):2748-58. doi: 10.1016/j.yexcr.2011.08.010. Epub 2011 Aug 22. Exp Cell Res. 2011. PMID: 21884696 Review.
-
Modeling Wnt signaling by CRISPR-Cas9 genome editing recapitulates neoplasia in human Barrett epithelial organoids.Cancer Lett. 2018 Nov 1;436:109-118. doi: 10.1016/j.canlet.2018.08.017. Epub 2018 Aug 23. Cancer Lett. 2018. PMID: 30144514 Free PMC article.
-
Role of adenomatous polyposis coli (APC) gene mutations in the pathogenesis of colorectal cancer; current status and perspectives.Biochimie. 2019 Feb;157:64-71. doi: 10.1016/j.biochi.2018.11.003. Epub 2018 Nov 8. Biochimie. 2019. PMID: 30414835 Review.
Cited by
-
Spatiotemporal Gradient and Instability of Wnt Induce Heterogeneous Growth and Differentiation of Human Intestinal Organoids.iScience. 2020 Aug 21;23(8):101372. doi: 10.1016/j.isci.2020.101372. Epub 2020 Jul 16. iScience. 2020. PMID: 32745985 Free PMC article.
-
CRISPR/Cas9: a powerful tool in colorectal cancer research.J Exp Clin Cancer Res. 2023 Nov 22;42(1):308. doi: 10.1186/s13046-023-02901-z. J Exp Clin Cancer Res. 2023. PMID: 37993945 Free PMC article. Review.
-
Immuno-genomic classification of colorectal cancer organoids reveals cancer cells with intrinsic immunogenic properties associated with patient survival.J Exp Clin Cancer Res. 2021 Jul 13;40(1):230. doi: 10.1186/s13046-021-02034-1. J Exp Clin Cancer Res. 2021. PMID: 34256801 Free PMC article.
-
Predicting colorectal cancer risk in FAP patients using patient-specific organoids.Cancer Gene Ther. 2025 Sep;32(9):997-1007. doi: 10.1038/s41417-025-00923-7. Epub 2025 Jul 22. Cancer Gene Ther. 2025. PMID: 40695959
-
Molecular subtypes of colorectal cancer in the era of precision oncotherapy: Current inspirations and future challenges.Cancer Med. 2024 Jul;13(14):e70041. doi: 10.1002/cam4.70041. Cancer Med. 2024. PMID: 39054866 Free PMC article. Review.
References
-
- Aoki R., Shoshkes-Carmel M., Gao N., Shin S., May C.L., Golson M.L., Zahm A.M., Ray M., Wiser C.L., Wright C.V.E., and Kaestner K.H.. 2016. Foxl1-expressing mesenchymal cells constitute the intestinal stem cell niche. Cell. Mol. Gastroenterol. Hepatol. 2:175–188. 10.1016/j.jcmgh.2015.12.004 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

