PML is recruited to heterochromatin during S phase and represses DAXX-mediated histone H3.3 chromatin assembly
- PMID: 30796101
- PMCID: PMC6451418
- DOI: 10.1242/jcs.220970
PML is recruited to heterochromatin during S phase and represses DAXX-mediated histone H3.3 chromatin assembly
Abstract
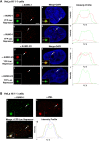
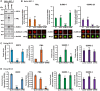
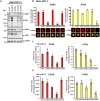
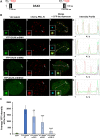
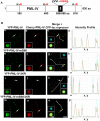
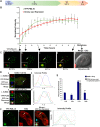
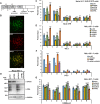
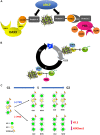
The incorporation of the histone H3 variant, H3.3, into chromatin by the H3.3-specific chaperone DAXX and the ATP-dependent chromatin remodeling factor ATRX is a critical mechanism for silencing repetitive DNA. DAXX and ATRX are also components of promyelocytic nuclear bodies (PML-NBs), which have been identified as sites of H3.3 chromatin assembly. Here, we use a transgene array that can be visualized in single living cells to investigate the mechanisms that recruit PML-NB proteins (i.e. PML, DAXX, ATRX, and SUMO-1, SUMO-2 and SUMO-3) to heterochromatin and their functions in H3.3 chromatin assembly. We show that DAXX and PML are recruited to the array through distinct SUMOylation-dependent mechanisms. Additionally, PML is recruited during S phase and its depletion increases H3.3 deposition. Since this effect is abrogated when PML and DAXX are co-depleted, it is likely that PML represses DAXX-mediated H3.3 chromatin assembly. Taken together, these results suggest that, at heterochromatin, PML-NBs coordinate H3.3 chromatin assembly with DNA replication, which has important implications for understanding how transcriptional silencing is established and maintained.
Keywords: ATRX; DAXX; Histone H3.3; PML; PML nuclear body; SUMOylation.
© 2019. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures
Similar articles
-
The death-associated protein DAXX is a novel histone chaperone involved in the replication-independent deposition of H3.3.Genes Dev. 2010 Jun 15;24(12):1253-65. doi: 10.1101/gad.566910. Epub 2010 May 26. Genes Dev. 2010. PMID: 20504901 Free PMC article.
-
Dynamics of histone H3.3 deposition in proliferating and senescent cells reveals a DAXX-dependent targeting to PML-NBs important for pericentromeric heterochromatin organization.Cell Cycle. 2014;13(2):249-67. doi: 10.4161/cc.26988. Epub 2013 Nov 5. Cell Cycle. 2014. PMID: 24200965 Free PMC article.
-
Single-cell analysis of Daxx and ATRX-dependent transcriptional repression.J Cell Sci. 2012 Nov 15;125(Pt 22):5489-501. doi: 10.1242/jcs.110148. Epub 2012 Sep 12. J Cell Sci. 2012. PMID: 22976303 Free PMC article.
-
Modulation of H3.3 chromatin assembly by PML: A way to regulate epigenetic inheritance.Bioessays. 2021 Oct;43(10):e2100038. doi: 10.1002/bies.202100038. Epub 2021 Aug 22. Bioessays. 2021. PMID: 34423467 Review.
-
The Dynamic Regulation of Daxx-Mediated Transcriptional Inhibition by SUMO and PML NBs.Int J Mol Sci. 2025 Jul 12;26(14):6703. doi: 10.3390/ijms26146703. Int J Mol Sci. 2025. PMID: 40724953 Free PMC article. Review.
Cited by
-
Phase-Separated Subcellular Compartmentation and Related Human Diseases.Int J Mol Sci. 2022 May 14;23(10):5491. doi: 10.3390/ijms23105491. Int J Mol Sci. 2022. PMID: 35628304 Free PMC article. Review.
-
DAXX Is a Crucial Factor for Proper Development of Mammalian Oocytes and Early Embryos.Int J Mol Sci. 2021 Jan 28;22(3):1313. doi: 10.3390/ijms22031313. Int J Mol Sci. 2021. PMID: 33525665 Free PMC article. Review.
-
PML modulates epigenetic composition of chromatin to regulate expression of pro-metastatic genes in triple-negative breast cancer.Nucleic Acids Res. 2023 Nov 10;51(20):11024-11039. doi: 10.1093/nar/gkad819. Nucleic Acids Res. 2023. PMID: 37823593 Free PMC article.
-
PML nuclear bodies and chromatin dynamics: catch me if you can!Nucleic Acids Res. 2020 Dec 2;48(21):11890-11912. doi: 10.1093/nar/gkaa828. Nucleic Acids Res. 2020. PMID: 33068409 Free PMC article. Review.
-
Nuclear body phase separation drives telomere clustering in ALT cancer cells.Mol Biol Cell. 2020 Aug 15;31(18):2048-2056. doi: 10.1091/mbc.E19-10-0589. Epub 2020 Jun 24. Mol Biol Cell. 2020. PMID: 32579423 Free PMC article.
References
-
- Banaszynski L. A., Wen D., Dewell S., Whitcomb S. J., Lin M., Diaz N., Elsässer S. J., Chapgier A., Goldberg A. D., Canaani E. et al. (2013). Hira-dependent histone H3.3 deposition facilitates PRC2 recruitment at developmental loci in ES cells. Cell 155, 107-120. 10.1016/j.cell.2013.08.061 - DOI - PMC - PubMed
-
- Bérubé N. G., Mangelsdorf M., Jagla M., Vanderluit J., Garrick D., Gibbons R. J., Higgs D. R., Slack R. S. and Picketts D. J. (2005). The chromatin-remodeling protein ATRX is critical for neuronal survival during corticogenesis. J. Clin. Invest. 115, 258-267. 10.1172/JCI200522329 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

