The C-terminal region of Net1 is an activator of RNA polymerase I transcription with conserved features from yeast to human
- PMID: 30802237
- PMCID: PMC6415870
- DOI: 10.1371/journal.pgen.1008006
The C-terminal region of Net1 is an activator of RNA polymerase I transcription with conserved features from yeast to human
Abstract
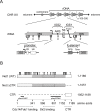
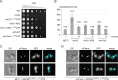
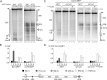
RNA polymerase I (Pol I) synthesizes ribosomal RNA (rRNA) in all eukaryotes, accounting for the major part of transcriptional activity in proliferating cells. Although basal Pol I transcription factors have been characterized in diverse organisms, the molecular basis of the robust rRNA production in vivo remains largely unknown. In S. cerevisiae, the multifunctional Net1 protein was reported to stimulate Pol I transcription. We found that the Pol I-stimulating function can be attributed to the very C-terminal region (CTR) of Net1. The CTR was required for normal cell growth and Pol I recruitment to rRNA genes in vivo and sufficient to promote Pol I transcription in vitro. Similarity with the acidic tail region of mammalian Pol I transcription factor UBF, which could partly functionally substitute for the CTR, suggests conserved roles for CTR-like domains in Pol I transcription from yeast to human.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
-
- Warner JR. The economics of ribosome biosynthesis in yeast. Trends Biochem Sci. 1999. November;24(11):437–40. - PubMed
-
- Boukhgalter B, Liu M, Guo A, Tripp M, Tran K, Huynh C, et al. Characterization of a fission yeast subunit of an RNA polymerase I essential transcription initiation factor, SpRrn7h/TAF(I)68, that bridges yeast and mammals: association with SpRrn11h and the core ribosomal RNA gene promoter. Gene. 2002. May 29;291(1–2):187–201. - PubMed
-
- Keys DA, Vu L, Steffan JS, Dodd JA, Yamamoto RT, Nogi Y, et al. RRN6 and RRN7 encode subunits of a multiprotein complex essential for the initiation of rDNA transcription by RNA polymerase I in Saccharomyces cerevisiae. Genes Dev. 1994. October 1;8(19):2349–62. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

