Hybridization is a recurrent evolutionary stimulus in wild yeast speciation
- PMID: 30804385
- PMCID: PMC6389940
- DOI: 10.1038/s41467-019-08809-7
Hybridization is a recurrent evolutionary stimulus in wild yeast speciation
Erratum in
-
Author Correction: Hybridization is a recurrent evolutionary stimulus in wild yeast speciation.Nat Commun. 2019 May 13;10(1):2199. doi: 10.1038/s41467-019-09702-z. Nat Commun. 2019. PMID: 31086180 Free PMC article.
Abstract
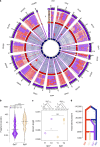
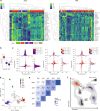
Hybridization can result in reproductively isolated and phenotypically distinct lineages that evolve as independent hybrid species. How frequently hybridization leads to speciation remains largely unknown. Here we examine the potential recurrence of hybrid speciation in the wild yeast Saccharomyces paradoxus in North America, which comprises two endemic lineages SpB and SpC, and an incipient hybrid species, SpC*. Using whole-genome sequences from more than 300 strains, we uncover the hybrid origin of another group, SpD, that emerged from hybridization between SpC* and one of its parental species, the widespread SpB. We show that SpD has the potential to evolve as a novel hybrid species, because it displays phenotypic novelties that include an intermediate transcriptome profile, and partial reproductive isolation with its most abundant sympatric parental species, SpB. Our findings show that repetitive cycles of divergence and hybridization quickly generate diversity and reproductive isolation, providing the raw material for speciation by hybridization.
Conflict of interest statement
The authors declare no competing interests.
Figures


Similar articles
-
Mitochondrial Recombination and Introgression during Speciation by Hybridization.Mol Biol Evol. 2017 Aug 1;34(8):1947-1959. doi: 10.1093/molbev/msx139. Mol Biol Evol. 2017. PMID: 28444332 Free PMC article.
-
Chromosomal variation segregates within incipient species and correlates with reproductive isolation.Mol Ecol. 2014 Sep;23(17):4362-72. doi: 10.1111/mec.12864. Epub 2014 Aug 20. Mol Ecol. 2014. PMID: 25039979
-
Speciation driven by hybridization and chromosomal plasticity in a wild yeast.Nat Microbiol. 2016 Jan 11;1:15003. doi: 10.1038/nmicrobiol.2015.3. Nat Microbiol. 2016. PMID: 27571751
-
Hybridization and the origin of new yeast lineages.FEMS Yeast Res. 2020 Aug 1;20(5):foaa040. doi: 10.1093/femsyr/foaa040. FEMS Yeast Res. 2020. PMID: 32658267 Free PMC article. Review.
-
The molecular evolutionary basis of species formation.Nat Rev Genet. 2010 Mar;11(3):175-80. doi: 10.1038/nrg2718. Epub 2010 Jan 6. Nat Rev Genet. 2010. PMID: 20051985 Review.
Cited by
-
The genomic landscape of transposable elements in yeast hybrids is shaped by structural variation and genotype-specific modulation of transposition rate.Elife. 2024 Feb 27;12:RP89277. doi: 10.7554/eLife.89277. Elife. 2024. PMID: 38411604 Free PMC article.
-
Similarities in biological processes can be used to bridge ecology and molecular biology.Evol Appl. 2020 Apr 13;13(6):1335-1350. doi: 10.1111/eva.12961. eCollection 2020 Jul. Evol Appl. 2020. PMID: 32684962 Free PMC article.
-
Interspecific Gene Exchange Introduces High Genetic Variability in Crop Pathogen.Genome Biol Evol. 2019 Nov 1;11(11):3095-3105. doi: 10.1093/gbe/evz224. Genome Biol Evol. 2019. PMID: 31603209 Free PMC article.
-
Genome structure reveals the diversity of mating mechanisms in Saccharomyces cerevisiae x Saccharomyces kudriavzevii hybrids, and the genomic instability that promotes phenotypic diversity.Microb Genom. 2020 Mar;6(3):e000333. doi: 10.1099/mgen.0.000333. Microb Genom. 2020. PMID: 32065577 Free PMC article.
-
Horizontal Transfer and Recombination Fuel Ty4 Retrotransposon Evolution in Saccharomyces.Genome Biol Evol. 2025 Jan 6;17(1):evaf004. doi: 10.1093/gbe/evaf004. Genome Biol Evol. 2025. PMID: 39786570 Free PMC article.
References
-
- Anderson E, Stebbins GL. Hybridization as an evolutionary stimulus. Evolution. 1954;8:378–388. doi: 10.1111/j.1558-5646.1954.tb01504.x. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

