Comparative Metagenomics and Network Analyses Provide Novel Insights Into the Scope and Distribution of β-Lactamase Homologs in the Environment
- PMID: 30804916
- PMCID: PMC6378392
- DOI: 10.3389/fmicb.2019.00146
Comparative Metagenomics and Network Analyses Provide Novel Insights Into the Scope and Distribution of β-Lactamase Homologs in the Environment
Abstract
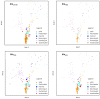
The β-lactams are the largest group of clinically applied antibiotics, and resistance to these is primarily associated with β-lactamases. There is increasing understanding that these enzymes are ubiquitous in natural environments and henceforth, elucidating the global diversity, distribution, and mobility of β-lactamase-encoding genes is crucial for holistically understanding resistance to these antibiotics. In this study, we screened 232 shotgun metagenomes from ten different environments against a custom-designed β-lactamase database, and subsequently analyzed β-lactamase homologs with a suite of bioinformatic platforms including cluster and network analyses. Three interrelated β-lactamase clusters encompassed all of the human and bovine feces metagenomes, while β-lactamases from soil, freshwater, glacier, marine, and wastewater grouped within a separate "environmental" cluster that displayed high levels of inter-network connectivity. Interestingly, almost no connectivity occurred between the "feces" and "environmental" clusters. We attributed this in part to the divergence in microbial community composition (dominance of Bacteroidetes and Firmicutes vs. Proteobacteria, respectively). The β-lactamase diversity in the "environmental" cluster was significantly higher than in human and bovine feces microbiomes. Several class A, B, C, and D β-lactamase homologs (bla CTX-M, bla KPC, bla GES) were ubiquitous in the "environmental" cluster, whereas bovine and human feces metagenomes were dominated by class A (primarily cfxA) β-lactamases. Collectively, this study highlights the ubiquitous presence and broad diversity of β-lactamase gene precursors in non-clinical environments. Furthermore, it suggests that horizontal transfer of β-lactamases to human-associated bacteria may be more plausible from animals than from terrestrial and aquatic microbes, seemingly due to phylogenetic similarities.
Keywords: antibiotic resistance; environment; metagenomics; network analysis; β-lactamases.
Figures
Similar articles
-
Distribution and Classification of Serine β-Lactamases in Brazilian Hospital Sewage and Other Environmental Metagenomes Deposited in Public Databases.Front Microbiol. 2016 Nov 15;7:1790. doi: 10.3389/fmicb.2016.01790. eCollection 2016. Front Microbiol. 2016. PMID: 27895627 Free PMC article.
-
Aeromonas spp. simultaneously harbouring bla(CTX-M-15), bla(SHV-12), bla(PER-1) and bla(FOX-2), in wild-growing Mediterranean mussel (Mytilus galloprovincialis) from Adriatic Sea, Croatia.Int J Food Microbiol. 2013 Sep 2;166(2):301-8. doi: 10.1016/j.ijfoodmicro.2013.07.010. Epub 2013 Jul 18. Int J Food Microbiol. 2013. PMID: 23973842
-
Detection of clinically important β-lactamases in commensal Escherichia coli of human and swine origin in western China.J Med Microbiol. 2012 Feb;61(Pt 2):233-238. doi: 10.1099/jmm.0.036806-0. Epub 2011 Sep 22. J Med Microbiol. 2012. PMID: 21940649
-
Resistance to cephalosporins and carbapenems in Gram-negative bacterial pathogens.Int J Med Microbiol. 2010 Aug;300(6):371-9. doi: 10.1016/j.ijmm.2010.04.005. Epub 2010 May 27. Int J Med Microbiol. 2010. PMID: 20537585 Review.
-
Beta-lactamase inhibitors: the story so far.Curr Med Chem. 2009;16(28):3740-65. doi: 10.2174/092986709789104957. Curr Med Chem. 2009. PMID: 19747143 Review.
Cited by
-
Antimicrobial Resistance of Non-O157 Shiga Toxin-Producing Escherichia coli Isolated from Humans and Domestic Animals.Antibiotics (Basel). 2021 Jan 14;10(1):74. doi: 10.3390/antibiotics10010074. Antibiotics (Basel). 2021. PMID: 33466678 Free PMC article.
-
Metagenomic insights into the antibiotic resistomes of typical Chinese dairy farm environments.Front Microbiol. 2022 Sep 28;13:990272. doi: 10.3389/fmicb.2022.990272. eCollection 2022. Front Microbiol. 2022. PMID: 36246251 Free PMC article.
-
Detection of blaCTX-M and blaDHA genes in stool samples of healthy people: comparison of culture- and shotgun metagenomic-based approaches.Front Microbiol. 2023 Aug 31;14:1236208. doi: 10.3389/fmicb.2023.1236208. eCollection 2023. Front Microbiol. 2023. PMID: 37720151 Free PMC article.
-
Plastics and the microbiome: impacts and solutions.Environ Microbiome. 2021 Jan 20;16(1):2. doi: 10.1186/s40793-020-00371-w. Environ Microbiome. 2021. PMID: 33902756 Free PMC article. Review.
-
Combined effects of composting and antibiotic administration on cattle manure-borne antibiotic resistance genes.Microbiome. 2021 Apr 1;9(1):81. doi: 10.1186/s40168-021-01006-z. Microbiome. 2021. PMID: 33795006 Free PMC article.
References
LinkOut - more resources
Full Text Sources

