Laying the Foundation for Crassulacean Acid Metabolism (CAM) Biodesign: Expression of the C4 Metabolism Cycle Genes of CAM in Arabidopsis
- PMID: 30804970
- PMCID: PMC6378705
- DOI: 10.3389/fpls.2019.00101
Laying the Foundation for Crassulacean Acid Metabolism (CAM) Biodesign: Expression of the C4 Metabolism Cycle Genes of CAM in Arabidopsis
Abstract
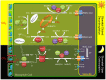
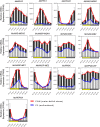
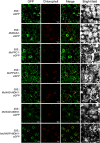
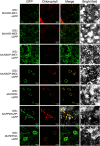
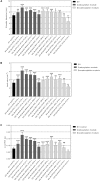
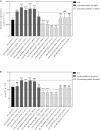
Crassulacean acid metabolism (CAM) is a specialized mode of photosynthesis that exploits a temporal CO2 pump with nocturnal CO2 uptake and concentration to reduce photorespiration, improve water-use efficiency (WUE), and optimize the adaptability of plants to hotter and drier climates. Introducing the CAM photosynthetic machinery into C3 (or C4) photosynthesis plants (CAM Biodesign) represents a potentially breakthrough strategy for improving WUE while maintaining high productivity. To optimize the success of CAM Biodesign approaches, the functional analysis of individual C4 metabolism cycle genes is necessary to identify the essential genes for robust CAM pathway introduction. Here, we isolated and analyzed the subcellular localizations of 13 enzymes and regulatory proteins of the C4 metabolism cycle of CAM from the common ice plant in stably transformed Arabidopsis thaliana. Six components of the carboxylation module were analyzed including beta-carbonic anhydrase (McBCA2), phosphoenolpyruvate carboxylase (McPEPC1), phosphoenolpyruvate carboxylase kinase (McPPCK1), NAD-dependent malate dehydrogenase (McNAD-MDH1, McNAD-MDH2), and NADP-dependent malate dehydrogenase (McNADP-MDH1). In addition, seven components of the decarboxylation module were analyzed including NAD-dependent malic enzyme (McNAD-ME1, McNAD-ME2), NADP-dependent malic enzyme (McNADP-ME1, NADP-ME2), pyruvate, orthophosphate dikinase (McPPDK), pyruvate, orthophosphate dikinase-regulatory protein (McPPDK-RP), and phosphoenolpyruvate carboxykinase (McPEPCK). Ectopic overexpression of most C4-metabolism cycle components resulted in increased rosette diameter, leaf area, and leaf fresh weight of A. thaliana except for McNADP-MDH1, McPPDK-RP, and McPEPCK. Overexpression of most carboxylation module components resulted in increased stomatal conductance and dawn/dusk titratable acidity (TA) as an indirect measure of organic acid (mainly malate) accumulation in A. thaliana. In contrast, overexpression of the decarboxylating malic enzymes reduced stomatal conductance and TA. This comprehensive study provides fundamental insights into the relative functional contributions of each of the individual components of the core C4-metabolism cycle of CAM and represents a critical first step in laying the foundation for CAM Biodesign.
Keywords: Arabidopsis thaliana; C4 metabolism; CAM biodesign; Mesembryanthemum crystallinum; crassulacean acid metabolism; ice plant; water-use efficiency.
Figures

Similar articles
-
Activity of enzymes of carbon metabolism during the induction of Crassulacean acid metabolism in Mesembryanthemum crystallinum L.Planta. 1982 Jun;155(1):8-16. doi: 10.1007/BF00402925. Planta. 1982. PMID: 24271620
-
Photosynthesis-related characteristics of the midrib and the interveinal lamina in leaves of the C3-CAM intermediate plant Mesembryanthemum crystallinum.Ann Bot. 2016 Jun;117(7):1141-51. doi: 10.1093/aob/mcw049. Epub 2016 Apr 18. Ann Bot. 2016. PMID: 27091507 Free PMC article.
-
Role of C4 photosynthetic enzyme isoforms in C3 plants and their potential applications in improving agronomic traits in crops.Photosynth Res. 2022 Dec;154(3):233-258. doi: 10.1007/s11120-022-00978-9. Epub 2022 Oct 29. Photosynth Res. 2022. PMID: 36309625 Review.
-
Intracellular Localization of Enzymes of Carbon Metabolism in Mesembryanthemum crystallinum Exhibiting C(3) Photosynthetic Characteristics or Performing Crassulacean Acid Metabolism.Plant Physiol. 1982 Feb;69(2):300-7. doi: 10.1104/pp.69.2.300. Plant Physiol. 1982. PMID: 16662197 Free PMC article.
-
Understanding trait diversity associated with crassulacean acid metabolism (CAM).Curr Opin Plant Biol. 2019 Jun;49:74-85. doi: 10.1016/j.pbi.2019.06.004. Epub 2019 Jul 5. Curr Opin Plant Biol. 2019. PMID: 31284077 Review.
Cited by
-
The first released available genome of the common ice plant ( Mesembryanthemum crystallinum L.) extended the research region on salt tolerance, C 3-CAM photosynthetic conversion, and halophilism.F1000Res. 2024 Jan 4;12:448. doi: 10.12688/f1000research.129958.4. eCollection 2023. F1000Res. 2024. PMID: 38618020 Free PMC article.
-
Crassulacean acid metabolism (CAM) at the crossroads: a special issue to honour 50 years of CAM research by Klaus Winter.Ann Bot. 2023 Nov 25;132(4):553-561. doi: 10.1093/aob/mcad160. Ann Bot. 2023. PMID: 37856823 Free PMC article. No abstract available.
-
Biosystems Design to Accelerate C3-to-CAM Progression.Biodes Res. 2020 Oct 10;2020:3686791. doi: 10.34133/2020/3686791. eCollection 2020. Biodes Res. 2020. PMID: 37849902 Free PMC article. Review.
-
Insights into the regulation of energy metabolism during the seed-to-seedling transition in marine angiosperm Zostera marina L.: Integrated metabolomic and transcriptomic analysis.Front Plant Sci. 2023 Mar 10;14:1130292. doi: 10.3389/fpls.2023.1130292. eCollection 2023. Front Plant Sci. 2023. PMID: 36968358 Free PMC article.
-
Undervalued potential of crassulacean acid metabolism for current and future agricultural production.J Exp Bot. 2019 Nov 29;70(22):6521-6537. doi: 10.1093/jxb/erz223. J Exp Bot. 2019. PMID: 31087091 Free PMC article. Review.
References
-
- Araújo W. L., Nunes-Nesi A., Osorio S., Usadel B., Fuentes D., Nagy R., et al. (2011b). Antisense inhibition of the iron-sulphur subunit of succinate dehydrogenase enhances photosynthesis and growth in tomato via an organic acid-mediated effect on stomatal aperture. Plant Cell 23 600–627. 10.1105/tpc.110.081224 - DOI - PMC - PubMed
-
- Astley H. M., Parsley K., Aubry S., Chastain C. J., Burnell J. N., Webb M. E., et al. (2011). The pyruvate, orthophosphate dikinase regulatory proteins of Arabidopsis are both bifunctional and interact with the catalytic and nucleotide-binding domains of pyruvate, orthophosphate dikinase. Plant J. 68 1070–1080. 10.1111/j.1365-313X.2011.04759.x - DOI - PubMed
-
- Backhausen J. E., Emmerlich A., Holtgrefe S., Horton P., Nast G., Rogers J. J., et al. (1998). Transgenic potato plants with altered expression levels of chloroplast NADP-malate dehydrogenase: interactions between photosynthetic electron transport and malate metabolism in leaves and in isolated intact chloroplasts. Planta 207 105–114. 10.1007/s004250050461 - DOI
-
- Badia M. B., Arias C. L., Tronconi M. A., Maurino V. G., Andreo C. S., Drincovich M. F., et al. (2015). Enhanced cytosolic NADP-ME2 activity in A. thaliana affects plant development, stress tolerance and specific diurnal and nocturnal cellular processes. Plant Sci. 240 193–203. 10.1016/j.plantsci.2015.09.015 - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

