Transcriptional Dynamics of Grain Development in Barley (Hordeum vulgare L.)
- PMID: 30813307
- PMCID: PMC6412674
- DOI: 10.3390/ijms20040962
Transcriptional Dynamics of Grain Development in Barley (Hordeum vulgare L.)
Abstract
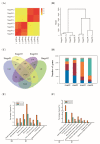
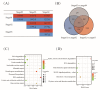
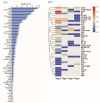
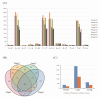
Grain development, as a vital process in the crop's life cycle, is crucial for determining crop quality and yield. However, the molecular basis and regulatory network of barley grain development is not well understood at present. Here, we investigated the transcriptional dynamics of barley grain development through RNA sequencing at four developmental phases, including early prestorage phase (3 days post anthesis (DPA)), late prestorage or transition phase (8 DPA), early storage phase (13 DPA), and levels off stages (18 DPA). Transcriptome profiling found that pronounced shifts occurred in the abundance of transcripts involved in both primary and secondary metabolism during grain development. The transcripts' activity was decreased during maturation while the largest divergence was observed between the transitions from prestorage phase to storage phase, which coincided with the physiological changes. Furthermore, the transcription factors, hormone signal transduction-related as well as sugar-metabolism-related genes, were found to play a crucial role in barley grain development. Finally, 4771 RNA editing events were identified in these four development stages, and most of the RNA editing genes were preferentially expressed at the prestore stage rather than in the store stage, which was significantly enriched in "essential" genes and plant hormone signal transduction pathway. These results suggested that RNA editing might act as a 'regulator' to control grain development. This study systematically dissected the gene expression atlas of barley grain development through transcriptome analysis, which not only provided the potential targets for further functional studies, but also provided insights into the dynamics of gene regulation underlying grain development in barley and beyond.
Keywords: Barley; Grain development; RNA editing; RNA-seq; Transcriptional dynamics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
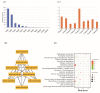
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

