A high-throughput SNP discovery strategy for RNA-seq data
- PMID: 30813897
- PMCID: PMC6391812
- DOI: 10.1186/s12864-019-5533-4
A high-throughput SNP discovery strategy for RNA-seq data
Abstract
Background: Single nucleotide polymorphisms (SNP) have been applied as important molecular markers in genetics and breeding studies. The rapid advance of next generation sequencing (NGS) provides a high-throughput means of SNP discovery. However, SNP development is limited by the availability of reliable SNP discovery methods. Especially, the optimum assembler and SNP caller for accurate SNP prediction from next generation sequencing data are not known.
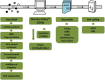
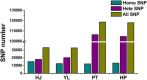
Results: Herein we performed SNP prediction based on RNA-seq data of peach and mandarin peel tissue under a comprehensive comparison of two paired-end read lengths (125 bp and 150 bp), five assemblers (Trinity, IDBA, oases, SOAPdenovo, Trans-abyss) and two SNP callers (GATK and GBS). The predicted SNPs were compared with the authentic SNPs identified via PCR amplification followed by gene cloning and sequencing procedures. A total of 40 and 240 authentic SNPs were presented in five anthocyanin biosynthesis related genes in peach and in nine carotenogenic genes in mandarin. Putative SNPs predicted from the same RNA-seq data with different strategies led to quite divergent results. The rate of false positive SNPs was significantly lower when the paired-end read length was 150 bp compared with 125 bp. Trinity was superior to the other four assemblers and GATK was substantially superior to GBS due to a low rate of missing authentic SNPs. The combination of assembler Trinity, SNP caller GATK, and the paired-end read length 150 bp had the best performance in SNP discovery with 100% accuracy both in peach and in mandarin cases. This strategy was applied to the characterization of SNPs in peach and mandarin transcriptomes.
Conclusions: Through comparison of authentic SNPs obtained by PCR cloning strategy and putative SNPs predicted from different combinations of five assemblers, two SNP callers, and two paired-end read lengths, we provided a reliable and efficient strategy, Trinity-GATK with 150 bp paired-end read length, for SNP discovery from RNA-seq data. This strategy discovered SNP at 100% accuracy in peach and mandarin cases and might be applicable to a wide range of plants and other organisms.
Keywords: GATK; Paired-end read length; RNA-seq; Single nucleotide polymorphism (SNP); Trinity.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Single nucleotide polymorphism discovery in bovine liver using RNA-seq technology.PLoS One. 2017 Feb 24;12(2):e0172687. doi: 10.1371/journal.pone.0172687. eCollection 2017. PLoS One. 2017. PMID: 28234981 Free PMC article.
-
Single Nucleotide Polymorphism Discovery in Bovine Pituitary Gland Using RNA-Seq Technology.PLoS One. 2016 Sep 8;11(9):e0161370. doi: 10.1371/journal.pone.0161370. eCollection 2016. PLoS One. 2016. PMID: 27606429 Free PMC article.
-
Gene-based SNP identification and validation in soybean using next-generation transcriptome sequencing.Mol Genet Genomics. 2018 Jun;293(3):623-633. doi: 10.1007/s00438-017-1410-5. Epub 2017 Dec 27. Mol Genet Genomics. 2018. PMID: 29280001
-
Normalization for Single-Cell RNA-Seq Data Analysis.Methods Mol Biol. 2019;1935:11-23. doi: 10.1007/978-1-4939-9057-3_2. Methods Mol Biol. 2019. PMID: 30758817 Review.
-
Genes, behavior and next-generation RNA sequencing.Genes Brain Behav. 2013 Feb;12(1):1-12. doi: 10.1111/gbb.12007. Epub 2012 Dec 28. Genes Brain Behav. 2013. PMID: 23194347 Free PMC article. Review.
Cited by
-
Whole Transcriptome Sequencing Unveils the Genomic Determinants of Putative Somaclonal Variation in Mint (Mentha L.).Int J Mol Sci. 2022 May 10;23(10):5291. doi: 10.3390/ijms23105291. Int J Mol Sci. 2022. PMID: 35628103 Free PMC article.
-
Modified "Allele-Specific qPCR" Method for SNP Genotyping Based on FRET.Front Plant Sci. 2022 Jan 10;12:747886. doi: 10.3389/fpls.2021.747886. eCollection 2021. Front Plant Sci. 2022. PMID: 35082803 Free PMC article.
-
Molecular targets and strategies in the development of nucleic acid cancer vaccines: from shared to personalized antigens.J Biomed Sci. 2024 Oct 9;31(1):94. doi: 10.1186/s12929-024-01082-x. J Biomed Sci. 2024. PMID: 39379923 Free PMC article. Review.
-
Development and application of the Faba_bean_130K targeted next-generation sequencing SNP genotyping platform based on transcriptome sequencing.Theor Appl Genet. 2021 Oct;134(10):3195-3207. doi: 10.1007/s00122-021-03885-0. Epub 2021 Jun 12. Theor Appl Genet. 2021. PMID: 34117907
-
Characterization of genome-wide genetic variations between two varieties of tea plant (Camellia sinensis) and development of InDel markers for genetic research.BMC Genomics. 2019 Dec 5;20(1):935. doi: 10.1186/s12864-019-6347-0. BMC Genomics. 2019. PMID: 31805860 Free PMC article.
References
-
- Jehan T, Lakhanpaul S. Single nucleotide polymorphism (SNP) – methods and applications in plant genetics: a review. Indian J Biotechnol. 2006;5:435–459.
-
- Hiremath PJ, Kumar A, Penmetsa RV, Farmer A, Schlueter JA, Chamarthi SK, Whaley AM, Carrasquilla-Garcia N, Gaur PM, Upadhyaya HD, et al. Large-scale development of cost-effective SNP marker assays for diversity assessment and genetic mapping in chickpea and comparative mapping in legumes. Plant Biotechnol J. 2012;10(6):1–17. doi: 10.1111/j.1467-7652.2012.00710.x. - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

