Metagenome-assembled genomes provide new insight into the microbial diversity of two thermal pools in Kamchatka, Russia
- PMID: 30816235
- PMCID: PMC6395817
- DOI: 10.1038/s41598-019-39576-6
Metagenome-assembled genomes provide new insight into the microbial diversity of two thermal pools in Kamchatka, Russia
Erratum in
-
Author Correction: Metagenome-assembled genomes provide new insight into the microbial diversity of two thermal pools in Kamchatka, Russia.Sci Rep. 2020 Feb 21;10(1):3454. doi: 10.1038/s41598-020-60390-y. Sci Rep. 2020. PMID: 32081994 Free PMC article.
Abstract
Culture-independent methods have contributed substantially to our understanding of global microbial diversity. Recently developed algorithms to construct whole genomes from environmental samples have further refined, corrected and revolutionized understanding of the tree of life. Here, we assembled draft metagenome-assembled genomes (MAGs) from environmental DNA extracted from two hot springs within an active volcanic ecosystem on the Kamchatka peninsula, Russia. This hydrothermal system has been intensively studied previously with regard to geochemistry, chemoautotrophy, microbial isolation, and microbial diversity. We assembled genomes of bacteria and archaea using DNA that had previously been characterized via 16S rRNA gene clone libraries. We recovered 36 MAGs, 29 of medium to high quality, and inferred their placement in a phylogenetic tree consisting of 3,240 publicly available microbial genomes. We highlight MAGs that were taxonomically assigned to groups previously underrepresented in available genome data. This includes several archaea (Korarchaeota, Bathyarchaeota and Aciduliprofundum) and one potentially new species within the bacterial genus Sulfurihydrogenibium. Putative functions in both pools were compared and are discussed in the context of their diverging geochemistry. This study adds comprehensive information about phylogenetic diversity and functional potential within two hot springs in the caldera of Kamchatka.
Conflict of interest statement
The authors declare no competing interests.
Figures



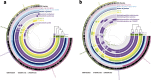
References
-
- Cowan, D., Tuffi, M., Mulako, I. & Cass, J. Terrestrial Hydrothermal Environments. In Life at Extremes: Environments, Organisms and Strategies for Survival (ed. Bell, E.) 1, 220–241 (CABI Press, London, UK, 2012).
-
- Beskrovnyy NS, et al. Presence of oil in hydrothermal systems associated with volcanism. Int. Geol. Rev. 1973;15:384–393. doi: 10.1080/00206817309475900. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

