An improved estimation of tRNA expression to better elucidate the coevolution between tRNA abundance and codon usage in bacteria
- PMID: 30816249
- PMCID: PMC6395768
- DOI: 10.1038/s41598-019-39369-x
An improved estimation of tRNA expression to better elucidate the coevolution between tRNA abundance and codon usage in bacteria
Abstract
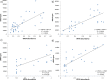
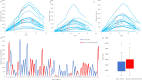
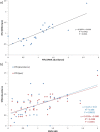
The degree to which codon usage can be explained by tRNA abundance in bacterial species is often inadequate, partly because differential tRNA abundance is often approximated by tRNA copy numbers. To better understand the coevolution between tRNA abundance and codon usage, we provide a better estimate of tRNA abundance by profiling tRNA mapped reads (tRNA tpm) using publicly available RNA Sequencing data. To emphasize the feasibility of our approach, we demonstrate that tRNA tpm is consistent with tRNA abundances derived from RNA fingerprinting experiments in Escherichia coli, Bacillus subtilis, and Salmonella enterica. Furthermore, we do not observe an appreciable reduction in tRNA sequencing efficiency due to post-transcriptional methylations in the seven bacteria studied. To determine optimal codons, we calculate codon usage in highly and lowly expressed genes determined by protein per transcript. We found that tRNA tpm is sensitive to identify more translationally optimal codons than gene copy number and early tRNA fingerprinting abundances. Additionally, tRNA tpm improves the predictive power of tRNA adaptation index over codon preference. Our results suggest that dependence of codon usage on tRNA availability is not always associated with species growth-rate. Conversely, tRNA availability is better optimized to codon usage in fast-growing than slow-growing species.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Evidence of multifaceted functions of codon usage in translation within the model beetle Tribolium castaneum.DNA Res. 2019 Dec 1;26(6):473-484. doi: 10.1093/dnares/dsz025. DNA Res. 2019. PMID: 31922535 Free PMC article.
-
Codon usage revisited: Lack of correlation between codon usage and the number of tRNA genes in enterobacteria.Biochem Biophys Res Commun. 2018 Aug 25;502(4):450-455. doi: 10.1016/j.bbrc.2018.05.168. Epub 2018 Jun 5. Biochem Biophys Res Commun. 2018. PMID: 29859934 Free PMC article.
-
Studies of codon usage and tRNA genes of 18 unicellular organisms and quantification of Bacillus subtilis tRNAs: gene expression level and species-specific diversity of codon usage based on multivariate analysis.Gene. 1999 Sep 30;238(1):143-55. doi: 10.1016/s0378-1119(99)00225-5. Gene. 1999. PMID: 10570992
-
Mechanisms governing codon usage bias and the implications for protein expression in the chloroplast of Chlamydomonas reinhardtii.Plant J. 2022 Nov;112(4):919-945. doi: 10.1111/tpj.15970. Epub 2022 Oct 19. Plant J. 2022. PMID: 36071273 Free PMC article. Review.
-
Preferential codon usage in prokaryotic genes: the optimal codon-anticodon interaction energy and the selective codon usage in efficiently expressed genes.Gene. 1982 Jun;18(3):199-209. doi: 10.1016/0378-1119(82)90157-3. Gene. 1982. PMID: 6751939 Review.
Cited by
-
The distinct translational landscapes of gram-negative Salmonella and gram-positive Listeria.Nat Commun. 2023 Dec 9;14(1):8167. doi: 10.1038/s41467-023-43759-1. Nat Commun. 2023. PMID: 38071303 Free PMC article.
-
Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon.Nat Commun. 2022 May 20;13(1):2815. doi: 10.1038/s41467-022-30390-9. Nat Commun. 2022. PMID: 35595777 Free PMC article.
-
Characterization of tRNA expression profiles in large offspring syndrome.BMC Genomics. 2022 Apr 7;23(1):273. doi: 10.1186/s12864-022-08496-7. BMC Genomics. 2022. PMID: 35392796 Free PMC article.
-
Plasma cell-free RNA characteristics in COVID-19 patients.Genome Res. 2022 Feb;32(2):228-241. doi: 10.1101/gr.276175.121. Epub 2022 Jan 21. Genome Res. 2022. PMID: 35064006 Free PMC article.
-
Bioinformatic Analysis Reveals the Role of Translation Elongation Efficiency Optimisation in the Evolution of Ralstonia Genus.Biology (Basel). 2023 Oct 16;12(10):1338. doi: 10.3390/biology12101338. Biology (Basel). 2023. PMID: 37887048 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

