A five-gene signature predicts overall survival of patients with papillary renal cell carcinoma
- PMID: 30822312
- PMCID: PMC6396926
- DOI: 10.1371/journal.pone.0211491
A five-gene signature predicts overall survival of patients with papillary renal cell carcinoma
Abstract
Background: The present study aims to investigate the gene expression changes in papillary renal cell carcinoma(pRCC) and screen several genes and associated pathways of papillary renal cell carcinoma progression.
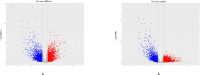
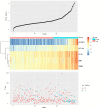
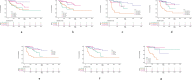
Methods: The papillary renal cell carcinoma RNA sequencing (RNA-seq) data set was downloaded from TCGA (The Cancer Genome Atlas). We identified the differentially expressed mRNAs between cancer and normal tissues and performed annotation of differentially expressed mRNAs to figure out the functions and pathways they were enriched in. Then, we constructed a risk score that relied on the 5-mRNA. The optimal value for the patients'classification risk level was identified by ROC analysis. The relationship between mRNA expression and prognosis of papillary renal cell carcinoma was evaluated by univariate Cox regression model. The 5-mRNA based risk score was validated in both complete set and testing set.
Result: In general, the 5-mRNA (CCNB2, IGF2BP3, KIF18A, PTTG1, and BUB1) were identified and validated, which can predict papillary renal cell carcinoma patient survival. This study revealed the 5-mRNA expression profile and the potential function of a single mRNA as a prognostic target for papillary renal cell carcinoma.
Conclusion: In addition, these findings may have significant implications for potential treatments options and prognosis for patients with papillary renal cell carcinoma.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

