Direct imaging of the recruitment and phosphorylation of S6K1 in the mTORC1 pathway in living cells
- PMID: 30833605
- PMCID: PMC6399282
- DOI: 10.1038/s41598-019-39410-z
Direct imaging of the recruitment and phosphorylation of S6K1 in the mTORC1 pathway in living cells
Abstract
Knowledge of protein signalling pathways in the working cell is seen as a primary route to identifying and developing targeted medicines. In recent years there has been a growing awareness of the importance of the mTOR pathway, making it an attractive target for therapeutic intervention in several diseases. Within this pathway we have focused on S6 kinase 1 (S6K1), the downstream phosphorylation substrate of mTORC1, and specifically identify its juxtaposition with mTORC1. When S6K1 is co-expressed with raptor we show that S6K1 is translocated from the nucleus to the cytoplasm. By developing a novel biosensor we demonstrate in real-time, that phosphorylation and de-phosphorylation of S6K1 occurs mainly in the cytoplasm of living cells. Furthermore, we show that the scaffold protein raptor, that typically recruits mTOR substrates, is not always involved in S6K1 phosphorylation. Overall, we demonstrate how FRET-FLIM imaging technology can be used to show localisation of S6K1 phosphorylation in living cells and hence a key site of action of inhibitors targeting mTOR phosphorylation.
Conflict of interest statement
Abdullah Ahmed has been funded by an iCASE PhD studentship BBSRC award supported by Evotec as the industrial partner. The award covers a stipend plus consumables and has no formal obligation or salaried links to Evotec. He declares no conflict of interest. All other authors declare no competing financial interest. We declare that we are in the process of allowing all plasmid mTORC1 constructs including SensOR to be made available through Cancer Research Technology Ximbio which provides a portal for the life science community to exchange knowledge and trade reagents. This site also provides repository for any accompanying raw data files.
Figures




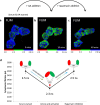

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

