Identifying Small Proteins by Ribosome Profiling with Stalled Initiation Complexes
- PMID: 30837344
- PMCID: PMC6401488
- DOI: 10.1128/mBio.02819-18
Identifying Small Proteins by Ribosome Profiling with Stalled Initiation Complexes
Abstract
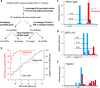
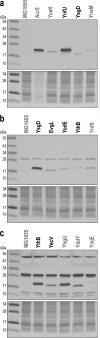
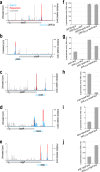
Small proteins consisting of 50 or fewer amino acids have been identified as regulators of larger proteins in bacteria and eukaryotes. Despite the importance of these molecules, the total number of small proteins remains unknown because conventional annotation pipelines usually exclude small open reading frames (smORFs). We previously identified several dozen small proteins in the model organism Escherichia coli using theoretical bioinformatic approaches based on sequence conservation and matches to canonical ribosome binding sites. Here, we present an empirical approach for discovering new proteins, taking advantage of recent advances in ribosome profiling in which antibiotics are used to trap newly initiated 70S ribosomes at start codons. This approach led to the identification of many novel initiation sites in intergenic regions in E. coli We tagged 41 smORFs on the chromosome and detected protein synthesis for all but three. Not only are the corresponding genes intergenic but they are also found antisense to other genes, in operons, and overlapping other open reading frames (ORFs), some impacting the translation of larger downstream genes. These results demonstrate the utility of this method for identifying new genes, regardless of their genomic context.IMPORTANCE Proteins comprised of 50 or fewer amino acids have been shown to interact with and modulate the functions of larger proteins in a range of organisms. Despite the possible importance of small proteins, the true prevalence and capabilities of these regulators remain unknown as the small size of the proteins places serious limitations on their identification, purification, and characterization. Here, we present a ribosome profiling approach with stalled initiation complexes that led to the identification of 38 new small proteins.
Keywords: Ribo-seq; alternate ORFs; antisense; genome annotation; leader peptide; small protein.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

