Application of Locked Nucleic Acid Oligonucleotides for siRNA Preclinical Bioanalytics
- PMID: 30837588
- PMCID: PMC6401054
- DOI: 10.1038/s41598-019-40187-4
Application of Locked Nucleic Acid Oligonucleotides for siRNA Preclinical Bioanalytics
Abstract
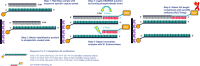
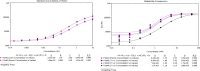
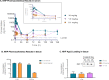
Despite the exquisite potential of siRNA as a therapeutic, the mechanism(s) responsible for the robust indirect exposure-response relationships have not been fully elucidated. To understand the siRNA properties linked to potent activity, requires the disposition of siRNA to be characterized. A technical challenge in the characterization is the detection and quantitation of siRNA from biological samples. Described herein, a Locked Nucleic Acid (LNA) Hybridization-Ligation ECL ELISA was designed for ultra-sensitive quantification of both sense and antisense strands of siRNA independent of structural modifica-tions. This assay was applied to measure siRNA in serum and tissue homogenate in preclinical species. We observed rapid clearance of siRNA from the systemic circulation which contrasted the prolonged accumulation within the tissue. The assay was also able to distinguish and quantify free siRNA from RNA-induced silencing complex (RISC) and Argonaute 2 (Ago2) associated with therapeutic siRNA. We utilized an orthogonal method, LC-MS, to investigate 3' exonuclease activity toward the antisense strand metabolism. Taken together, we have demonstrated that the LNA Hybridization-Ligation ECL ELISA is arobust analytical method with direct application to measuring the exposure of siRNA therapeutics seamlessly across biological matrices.
Conflict of interest statement
All authors of this manuscript are employees and shareholders of Amgen, Inc.
Figures

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

