Selection for novel metabolic capabilities in Salmonella enterica
- PMID: 30848832
- PMCID: PMC6593847
- DOI: 10.1111/evo.13713
Selection for novel metabolic capabilities in Salmonella enterica
Abstract
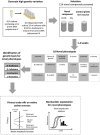
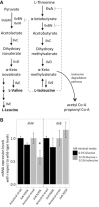
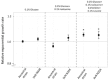
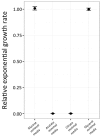
Bacteria are known to display extensive metabolic diversity and many studies have shown that they can use an extensive repertoire of small molecules as carbon- and energy sources. However, it is less clear to what extent a bacterium can expand its existing metabolic capabilities by acquiring mutations that, for example, rewire its metabolic pathways. To investigate this capability and potential for evolution of novel phenotypes, we sampled large populations of mutagenized Salmonella enterica to select very rare mutants that can grow on minimal media containing 124 low molecular weight compounds as sole carbon sources. We found mutants growing on 18 of these novel carbon sources, and identified the causal mutations that allowed growth for four of them. Mutations that relieve physiological constraints or increase expression of existing pathways were found to be important contributors to the novel phenotypes. For the remaining 14 novel phenotypes, whole genome sequencing of independent mutants and genetic analysis suggested that these novel metabolic phenotypes result from a combination of multiple mutations. This work, by virtue of identifying the genetic and mechanistic basis for new metabolic capabilities, sheds light on the properties of adaptive landscapes underlying the evolution of novel phenotypes.
Keywords: Carbon sources; experimental evolution; new metabolic functions; novel phenotypes.
© 2019 The Author(s). Evolution published by Wiley Periodicals, Inc. on behalf of The Society for the Study of Evolution.
Conflict of interest statement
The authors report no conflict of interest.
Figures

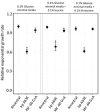
References
-
- Barberán, A. , Ramirez K. S., Leff J. W., Bradford M. A., Wall D. H., and Fierer N.. 2014. Why are some microbes more ubiquitous than others? Predicting the habitat breadth of soil bacteria. Ecol. Lett. 17:794–802. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
