Random Walk Enzymes: Information Theory, Quantum Isomorphism, and Entropy Dispersion
- PMID: 30848911
- PMCID: PMC6449183
- DOI: 10.1021/acs.jpca.9b00910
Random Walk Enzymes: Information Theory, Quantum Isomorphism, and Entropy Dispersion
Abstract
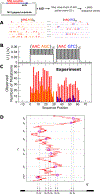
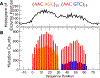
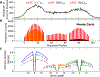
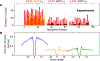
Activation-induced deoxycytidine deaminase (AID) is a key enzyme in the human immune system. AID binds to and catalyzes random point mutations on the immunoglobulin (Ig) gene, leading to diversification of the Ig gene sequence by random walk motions, scanning for cytidines and turning them to uracils. The mutation patterns deposited by AID on its substrate DNA sequences can be interpreted as random binary words, and the information content of this stochastically generated library of mutated DNA sequences can be measured by its entropy. In this paper, we derive an analytical formula for this entropy and show that the stochastic scanning + catalytic dynamics of AID is controlled by a characteristic length that depends on the diffusion coefficient of AID and the catalytic rate. Experiments showed that the deamination rates have a sequence context dependence, where mutations are generated at higher intensities on DNA sequences with higher densities of mutable sites. We derive an isomorphism between this classical system and a quantum mechanical model and use this isomorphism to explain why AID appears to focus its scanning on regions with higher concentrations of deaminable sites. Using path integral Monte Carlo simulations of the quantum isomorphic system, we demonstrate how AID's scanning indeed depends on the context of the DNA sequence and how this affects the entropy of the library of generated mutant clones. Examining detailed features in the entropy of the experimentally generated clone library, we provide clear evidence that the random walk of AID on its substrate DNA is focused near hot spots. The model calculations applied to the experimental data show that the observed per-site mutation frequencies display similar contextual dependences as observed in the experiments, in which hot motifs are located adjacent to several different types of hot and cold motifs.
Figures

References
-
- Muramatsu M; Kinoshita K; Fagarasan S; Yamada S; Shinkai Y; Honjo T, Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 2000, 102, 553–563. - PubMed
-
- Pham P; Bransteitter R; Petruska J; Goodman MF, Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature 2003, 424 (6944), 103–7. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

