A network-based approach to identify DNA methylation and its involved molecular pathways in testicular germ cell tumors
- PMID: 30854095
- PMCID: PMC6400810
- DOI: 10.7150/jca.27491
A network-based approach to identify DNA methylation and its involved molecular pathways in testicular germ cell tumors
Abstract
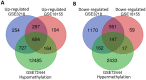
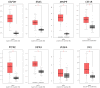
Background: Testicular germ cell tumors (TGCT) is the most common testicular malignancy threaten young male reproductive health. This study aimed to identify aberrantly methylated-differentially expressed genes and pathways in TGCT by comprehensive bioinformatics analysis. Methods: Data of gene expression microarrays (GSE3218, GSE18155) and gene methylation microarrays (GSE72444) were collected from GEO database. Integrated analysis acquired aberrantly methylated-genes. Functional and pathway enrichment analysis were performed using DAVID database. Protein-protein interaction (PPI) network was constructed by STRING and App Mcode was used for module analysis. GEPIA platform and DiseaseMeth database were used for confirming the expression and methylation levels of hub genes. Finally, Human Protein Atlas database was performed to evaluate the prognostic significance. Results: Totally 604 hypomethylation-high expression and 147 hypermethylation-low genes were identified. The high expressed genes were enriched in biological processes of cell proliferation and migration. The top 8 hub genes of PPI network were GAPDH, VEGFA, PTPRC, RIPK4, MMP9, CSF1R, KRAS and FN1. After validation in GEPIA platform, all hub genes were elevated in TGCT tissues. Only MMP9, CSF1R and PTPRC showed hypomethylation-high expression status, which predicted the poor outcome of TGCT patients. Conclusion: Our study indicated possible aberrantly methylated-differentially expressed genes and pathways in TGCT by bioinformatics analysis, which may provide novel insights for unraveling pathogenesis of TGCT.
Keywords: bioinformatics analysis; genetic etiology; methylation; testicular germ cell tumors.
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures




Similar articles
-
Aberrantly methylated-differentially expressed genes and pathways in colorectal cancer.Cancer Cell Int. 2017 Aug 7;17:75. doi: 10.1186/s12935-017-0444-4. eCollection 2017. Cancer Cell Int. 2017. PMID: 28794688 Free PMC article.
-
DNA methylation biomarkers for nasopharyngeal carcinoma.PLoS One. 2020 Apr 9;15(4):e0230524. doi: 10.1371/journal.pone.0230524. eCollection 2020. PLoS One. 2020. PMID: 32271791 Free PMC article.
-
Identification of aberrantly methylated differentially expressed genes in age-related macular degeneration.Medicine (Baltimore). 2019 Apr;98(14):e15083. doi: 10.1097/MD.0000000000015083. Medicine (Baltimore). 2019. PMID: 30946360 Free PMC article.
-
[Molecular Pathogenesis of Testicular Germ Cell Tumors].Klin Onkol. 2017 Winter;30(6):412-419. doi: 10.14735/amko2017412. Klin Onkol. 2017. PMID: 29271211 Review. Czech.
-
Recent Advancements in Research on DNA Methylation and Testicular Germ Cell Tumors: Unveiling the Intricate Relationship.Biomedicines. 2024 May 8;12(5):1041. doi: 10.3390/biomedicines12051041. Biomedicines. 2024. PMID: 38791003 Free PMC article. Review.
Cited by
-
Identification of potential core genes and miRNAs in testicular seminoma via bioinformatics analysis.Mol Med Rep. 2019 Nov;20(5):4013-4022. doi: 10.3892/mmr.2019.10684. Epub 2019 Sep 16. Mol Med Rep. 2019. PMID: 31545448 Free PMC article.
-
TNFSF10, an autophagy related gene, was a prognostic and immune infiltration marker in skin cutaneous melanoma.J Cancer. 2023 Jul 31;14(13):2417-2430. doi: 10.7150/jca.86735. eCollection 2023. J Cancer. 2023. PMID: 37670976 Free PMC article.
-
CSF1R methylation is a key regulatory mechanism of tumor-associated macrophages in hepatocellular carcinoma.Oncol Lett. 2020 Aug;20(2):1835-1845. doi: 10.3892/ol.2020.11726. Epub 2020 Jun 11. Oncol Lett. 2020. PMID: 32724427 Free PMC article.
-
Hypermethylation and global remodelling of DNA methylation is associated with acquired cisplatin resistance in testicular germ cell tumours.Epigenetics. 2021 Oct;16(10):1071-1084. doi: 10.1080/15592294.2020.1834926. Epub 2020 Oct 30. Epigenetics. 2021. PMID: 33126827 Free PMC article.
-
Between a Rock and a Hard Place: An Epigenetic-Centric View of Testicular Germ Cell Tumors.Cancers (Basel). 2021 Mar 25;13(7):1506. doi: 10.3390/cancers13071506. Cancers (Basel). 2021. PMID: 33805941 Free PMC article. Review.
References
-
- Manku G, Hueso A, Brimo F, Chan P, Gonzalez-Peramato P, Jabado N. et al. Changes in the expression profiles of claudins during gonocyte differentiation and in seminomas. Andrology. 2016;4:95–110. - PubMed
-
- International Prognostic Factors Study G, Lorch A, Beyer J, Bascoul-Mollevi C, Kramar A, Einhorn LH. et al. Prognostic factors in patients with metastatic germ cell tumors who experienced treatment failure with cisplatin-based first-line chemotherapy. J Clin Oncol. 2010;28:4906–11. - PubMed
-
- Turnbull C, Rahman N. Genome-wide association studies provide new insights into the genetic basis of testicular germ-cell tumour. Int J Androl. 2011;34:e86–96. discussion e-7. - PubMed
-
- Hanna NH, Einhorn LH. Testicular cancer-discoveries and updates. N Engl J Med. 2014;371:2005–16. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

