Profiling the Abiotic Stress Responsive microRNA Landscape of Arabidopsis thaliana
- PMID: 30857364
- PMCID: PMC6473545
- DOI: 10.3390/plants8030058
Profiling the Abiotic Stress Responsive microRNA Landscape of Arabidopsis thaliana
Abstract
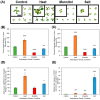
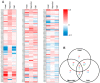
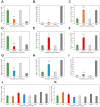
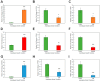
It is well established among interdisciplinary researchers that there is an urgent need to address the negative impacts that accompany climate change. One such negative impact is the increased prevalence of unfavorable environmental conditions that significantly contribute to reduced agricultural yield. Plant microRNAs (miRNAs) are key gene expression regulators that control development, defense against invading pathogens and adaptation to abiotic stress. Arabidopsis thaliana (Arabidopsis) can be readily molecularly manipulated, therefore offering an excellent experimental system to alter the profile of abiotic stress responsive miRNA/target gene expression modules to determine whether such modification enables Arabidopsis to express an altered abiotic stress response phenotype. Towards this goal, high throughput sequencing was used to profile the miRNA landscape of Arabidopsis whole seedlings exposed to heat, drought and salt stress, and identified 121, 123 and 118 miRNAs with a greater than 2-fold altered abundance, respectively. Quantitative reverse transcriptase polymerase chain reaction (RT-qPCR) was next employed to experimentally validate miRNA abundance fold changes, and to document reciprocal expression trends for the target genes of miRNAs determined abiotic stress responsive. RT-qPCR also demonstrated that each miRNA/target gene expression module determined to be abiotic stress responsive in Arabidopsis whole seedlings was reflective of altered miRNA/target gene abundance in Arabidopsis root and shoot tissues post salt stress exposure. Taken together, the data presented here offers an excellent starting platform to identify the miRNA/target gene expression modules for future molecular manipulation to generate plant lines that display an altered response phenotype to abiotic stress.
Keywords: Arabidopsis thaliana; RT-qPCR; abiotic stress; drought stress; heat stress; miRNA target gene expression; microRNAs (miRNAs); salt stress.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
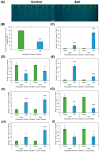
References
-
- Ramankutty N., Evan A.T., Monfreda C., Foley J.A. Farming the plant: 1. Geographic distribution of global agricultural lands in the year 2000. Glob. Biogeochem. Cycles. 2008:22. doi: 10.1029/2007GB002952. - DOI
LinkOut - more resources
Full Text Sources

