Analysis of error profiles in deep next-generation sequencing data
- PMID: 30867008
- PMCID: PMC6417284
- DOI: 10.1186/s13059-019-1659-6
Analysis of error profiles in deep next-generation sequencing data
Abstract
Background: Sequencing errors are key confounding factors for detecting low-frequency genetic variants that are important for cancer molecular diagnosis, treatment, and surveillance using deep next-generation sequencing (NGS). However, there is a lack of comprehensive understanding of errors introduced at various steps of a conventional NGS workflow, such as sample handling, library preparation, PCR enrichment, and sequencing. In this study, we use current NGS technology to systematically investigate these questions.
Results: By evaluating read-specific error distributions, we discover that the substitution error rate can be computationally suppressed to 10-5 to 10-4, which is 10- to 100-fold lower than generally considered achievable (10-3) in the current literature. We then quantify substitution errors attributable to sample handling, library preparation, enrichment PCR, and sequencing by using multiple deep sequencing datasets. We find that error rates differ by nucleotide substitution types, ranging from 10-5 for A>C/T>G, C>A/G>T, and C>G/G>C changes to 10-4 for A>G/T>C changes. Furthermore, C>T/G>A errors exhibit strong sequence context dependency, sample-specific effects dominate elevated C>A/G>T errors, and target-enrichment PCR led to ~ 6-fold increase of overall error rate. We also find that more than 70% of hotspot variants can be detected at 0.1 ~ 0.01% frequency with the current NGS technology by applying in silico error suppression.
Conclusions: We present the first comprehensive analysis of sequencing error sources in conventional NGS workflows. The error profiles revealed by our study highlight new directions for further improving NGS analysis accuracy both experimentally and computationally, ultimately enhancing the precision of deep sequencing.
Keywords: Deep sequencing; Detection; Error rate; Hotspot mutation; Subclonal; Substitution.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
A pending patent application has been filed based on the research disclosed in this manuscript; the patent does not restrict the research use of the findings in this article. The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures





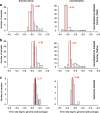

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

