Genomic plasticity associated with antimicrobial resistance in Vibrio cholerae
- PMID: 30867296
- PMCID: PMC6442563
- DOI: 10.1073/pnas.1900141116
Genomic plasticity associated with antimicrobial resistance in Vibrio cholerae
Abstract
The Bay of Bengal is known as the epicenter for seeding several devastating cholera outbreaks across the globe. Vibrio cholerae, the etiological agent of cholera, has extraordinary competency to acquire exogenous DNA by horizontal gene transfer (HGT) and adapt them into its genome for structuring metabolic processes, developing drug resistance, and colonizing the human intestine. Antimicrobial resistance (AMR) in V. cholerae has become a global concern. However, little is known about the identity of the resistance traits, source of AMR genes, acquisition process, and stability of the genetic elements linked with resistance genes in V. cholerae Here we present details of AMR profiles of 443 V. cholerae strains isolated from the stool samples of diarrheal patients from two regions of India. We sequenced the whole genome of multidrug-resistant (MDR) and extensively drug-resistant (XDR) V. cholerae to identify AMR genes and genomic elements that harbor the resistance traits. Our genomic findings were further confirmed by proteome analysis. We also engineered the genome of V. cholerae to monitor the importance of the autonomously replicating plasmid and core genome in the resistance profile. Our findings provided insights into the genomes of recent cholera isolates and identified several acquired traits including plasmids, transposons, integrative conjugative elements (ICEs), pathogenicity islands (PIs), prophages, and gene cassettes that confer fitness to the pathogen. The knowledge generated from this study would help in better understanding of V. cholerae evolution and management of cholera disease by providing clinical guidance on preferred treatment regimens.
Keywords: antimicrobial resistance; cholera; genome; mobile genetic elements; proteome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
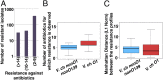






References
-
- Clemens JD, Nair GB, Ahmed T, Qadri F, Holmgren J. Cholera. Lancet. 2017;390:1539–1549. - PubMed
-
- Das B, et al. Molecular evolution and functional divergence of Vibrio cholerae. Curr Opin Infect Dis. 2016;29:520–527. - PubMed
-
- Meibom KL, Blokesch M, Dolganov NA, Wu CY, Schoolnik GK. Chitin induces natural competence in Vibrio cholerae. Science. 2005;310:1824–1827. - PubMed
-
- Kitaoka M, Miyata ST, Unterweger D, Pukatzki S. Antibiotic resistance mechanisms of Vibrio cholerae. J Med Microbiol. 2011;60:397–407. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

