Plasmid-Based CRISPR-Cas9 Gene Editing in Multiple Candida Species
- PMID: 30867327
- PMCID: PMC6416365
- DOI: 10.1128/mSphere.00125-19
Plasmid-Based CRISPR-Cas9 Gene Editing in Multiple Candida Species
Erratum in
-
Correction for Lombardi et al., "Plasmid-Based CRISPR-Cas9 Gene Editing in Multiple Candida Species".mSphere. 2020 Jun 10;5(3):e00494-20. doi: 10.1128/mSphere.00494-20. mSphere. 2020. PMID: 32522782 Free PMC article. No abstract available.
Abstract
Many Candida species that cause infection have diploid genomes and do not undergo classical meiosis. The application of clustered regularly interspaced short palindromic repeat-Cas9 (CRISPR-Cas9) gene editing systems has therefore greatly facilitated the generation of gene disruptions and the introduction of specific polymorphisms. However, CRISPR methods are not yet available for all Candida species. We describe here an adaption of a previously developed CRISPR system in Candida parapsilosis that uses an autonomously replicating plasmid. Guide RNAs can be introduced in a single cloning step and are released by cleavage between a tRNA and a ribozyme. The plasmid also contains CAS9 and a selectable nourseothricin SAT1 marker. It can be used for markerless editing in C. parapsilosis, C. orthopsilosis, and C. metapsilosis We also show that CRISPR can easily be used to introduce molecular barcodes and to reintroduce wild-type sequences into edited strains. Heterozygous mutations can be generated, either by careful selection of the distance between the polymorphism and the Cas9 cut site or by providing two different repair templates at the same time. In addition, we have constructed a different autonomously replicating plasmid for CRISPR-Cas9 editing in Candida tropicalis We show that editing can easily be carried out in multiple C. tropicalis isolates. Nonhomologous end joining (NHEJ) repair occurs at a high level in C. metapsilosis and C. tropicalisIMPORTANCECandida species are a major cause of infection worldwide. The species associated with infection vary with geographical location and with patient population. Infection with Candida tropicalis is particularly common in South America and Asia, and Candida parapsilosis infections are more common in the very young. Molecular methods for manipulating the genomes of these species are still lacking. We describe a simple and efficient CRISPR-based gene editing system that can be applied in the C. parapsilosis species group, including the sister species Candida orthopsilosis and Candida metapsilosis We have also constructed a separate system for gene editing in C. tropicalis.
Keywords: CRISPR; Candida; genome editing.
Copyright © 2019 Lombardi et al.
Figures
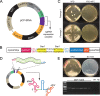




References
-
- Kurtzman C, Fell JW, Boekhout T. 2011. The yeasts: a taxonomic study. Elsevier, Philadelphia, PA.
-
- Pfaller MA, Pappas PG, Wingard JR. 2006. Invasive fungal pathogens: current epidemiological trends. Clin Infect Dis 43:S3–S14. doi: 10.1086/504490. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

