Hexamethylene bisacetamide impairs NK cell-mediated clearance of acute T lymphoblastic leukemia cells and HIV-1-infected T cells that exit viral latency
- PMID: 30867508
- PMCID: PMC6416400
- DOI: 10.1038/s41598-019-40760-x
Hexamethylene bisacetamide impairs NK cell-mediated clearance of acute T lymphoblastic leukemia cells and HIV-1-infected T cells that exit viral latency
Abstract
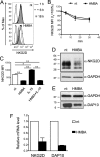
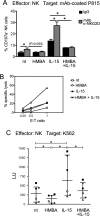
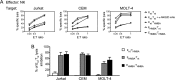
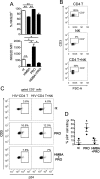
The hexamethylene bisacetamide (HMBA) anticancer drug was dismissed due to limited efficacy in leukemic patients but it may re-enter into the clinics in HIV-1 eradication strategies because of its recently disclosed capacity to reactivate latent virus. Here, we investigated the impact of HMBA on the cytotoxicity of natural killer (NK) cells against acute T lymphoblastic leukemia (T-ALL) cells or HIV-1-infected T cells that exit from latency. We show that in T-ALL cells HMBA upmodulated MICB and ULBP2 ligands for the NKG2D activating receptor. In a primary CD4+ T cell-based latency model, HMBA did not reactivate HIV-1, yet enhanced ULBP2 expression on cells harboring virus reactivated by prostratin (PRO). However, HMBA reduced the expression of NKG2D and its DAP10 adaptor in NK cells, hence impairing NKG2D-mediated cytotoxicity and DAP10-dependent response to IL-15 stimulation. Alongside, HMBA dampened killing of T-ALL targets by IL-15-activated NK cells and impaired NK cell-mediated clearance of PRO-reactivated HIV-1+ cells. Overall, our results demonstrate a dominant detrimental effect of HMBA on the NKG2D pathway that crucially controls NK cell-mediated killing of tumors and virus-infected cells, providing one possible explanation for poor clinical outcome in HMBA-treated cancer patients and raising concerns for future therapeutic application of this drug.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Potential of the NKG2D/NKG2DL Axis in NK Cell-Mediated Clearance of the HIV-1 Reservoir.Int J Mol Sci. 2019 Sep 11;20(18):4490. doi: 10.3390/ijms20184490. Int J Mol Sci. 2019. PMID: 31514330 Free PMC article. Review.
-
The histone deacetylase inhibitor SAHA simultaneously reactivates HIV-1 from latency and up-regulates NKG2D ligands sensitizing for natural killer cell cytotoxicity.Virology. 2017 Oct;510:9-21. doi: 10.1016/j.virol.2017.06.033. Epub 2017 Jul 6. Virology. 2017. PMID: 28689087
-
Combinations of Histone Deacetylase Inhibitors with Distinct Latency Reversing Agents Variably Affect HIV Reactivation and Susceptibility to NK Cell-Mediated Killing of T Cells That Exit Viral Latency.Int J Mol Sci. 2021 Jun 22;22(13):6654. doi: 10.3390/ijms22136654. Int J Mol Sci. 2021. PMID: 34206330 Free PMC article.
-
Soluble ligands for the NKG2D receptor are released during HIV-1 infection and impair NKG2D expression and cytotoxicity of NK cells.FASEB J. 2013 Jun;27(6):2440-50. doi: 10.1096/fj.12-223057. Epub 2013 Feb 8. FASEB J. 2013. PMID: 23395909
-
Release of Soluble Ligands for the Activating NKG2D Receptor: One More Immune Evasion Strategy Evolved by HIV-1 ?Curr Drug Targets. 2016;17(1):54-64. doi: 10.2174/1389450116666150630110329. Curr Drug Targets. 2016. PMID: 26122035 Review.
Cited by
-
Early ART initiation during infancy preserves natural killer cells in young European adolescents living with HIV (CARMA cohort).J Int AIDS Soc. 2021 Jul;24(7):e25717. doi: 10.1002/jia2.25717. J Int AIDS Soc. 2021. PMID: 34235857 Free PMC article.
-
Impact of IL-15 and latency reversing agent combinations in the reactivation and NK cell-mediated suppression of the HIV reservoir.Sci Rep. 2022 Nov 3;12(1):18567. doi: 10.1038/s41598-022-23010-5. Sci Rep. 2022. PMID: 36329160 Free PMC article.
-
Potential of the NKG2D/NKG2DL Axis in NK Cell-Mediated Clearance of the HIV-1 Reservoir.Int J Mol Sci. 2019 Sep 11;20(18):4490. doi: 10.3390/ijms20184490. Int J Mol Sci. 2019. PMID: 31514330 Free PMC article. Review.
-
Medicinal Chemistry of Anti-HIV-1 Latency Chemotherapeutics: Biotargets, Binding Modes and Structure-Activity Relationship Investigation.Molecules. 2022 Dec 20;28(1):3. doi: 10.3390/molecules28010003. Molecules. 2022. PMID: 36615199 Free PMC article. Review.
-
A functional mammalian display screen identifies rare antibodies that stimulate NK cell-mediated cytotoxicity.Proc Natl Acad Sci U S A. 2021 Aug 3;118(31):e2104099118. doi: 10.1073/pnas.2104099118. Proc Natl Acad Sci U S A. 2021. PMID: 34330834 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

