Genomic discovery of ion channel genes in the central nervous system of the lamprey Petromyzon marinus
- PMID: 30878501
- PMCID: PMC6579644
- DOI: 10.1016/j.margen.2019.03.003
Genomic discovery of ion channel genes in the central nervous system of the lamprey Petromyzon marinus
Abstract
The lamprey is a popular animal model for a number of types of neurobiology studies, including organization and operation of locomotor and respiratory systems, behavioral recovery following spinal cord injury (SCI), cellular and synaptic neurophysiology, comparative neuroanatomy, neuropharmacology, and neurodevelopment. Yet relatively little work has been done on the molecular underpinnings of nervous system function in lamprey. This is due in part to a paucity of gene information for some of the most fundamental proteins involved in neural activity: ion channels. We report here 47 putative ion channel sequences in the central nervous system (CNS) of larval lampreys from the predicted coding sequences (CDS) discovered in the P. marinus genome. These include 32 potassium (K+) channels, six sodium (Na+) channels, and nine calcium (Ca2+) channels. Through RT-PCR, we examined the distribution of these ion channels in the anterior (ARRN), middle (MRRN), and posterior (PRRN) rhombencephalic reticular nuclei, as well as the spinal cord (SC). This study lays the foundation for incorporating more advanced molecular techniques to investigate the role of ion channels in the neural networks of the lamprey.
Keywords: Genome; Ion channel; Reticulospinal.
Copyright © 2019 Elsevier B.V. All rights reserved.
Conflict of interest statement
Conflict of Interest Statement
The authors declare that they have no competing interests.
Figures

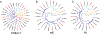
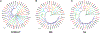

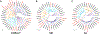

Similar articles
-
Differential effects of the reticulospinal system on locomotion in lamprey.J Neurophysiol. 1998 Jul;80(1):103-12. doi: 10.1152/jn.1998.80.1.103. J Neurophysiol. 1998. PMID: 9658032
-
Descending brain neurons in larval lamprey: spinal projection patterns and initiation of locomotion.Exp Neurol. 2010 Aug;224(2):527-41. doi: 10.1016/j.expneurol.2010.05.016. Epub 2010 May 25. Exp Neurol. 2010. PMID: 20510243 Free PMC article.
-
Inhibitory descending rhombencephalic projections in larval sea lamprey.Neuroscience. 2011 Oct 27;194:1-10. doi: 10.1016/j.neuroscience.2011.08.017. Epub 2011 Aug 12. Neuroscience. 2011. PMID: 21856380
-
Initiation of locomotion in lampreys.Brain Res Rev. 2008 Jan;57(1):172-82. doi: 10.1016/j.brainresrev.2007.07.016. Epub 2007 Aug 22. Brain Res Rev. 2008. PMID: 17916380 Review.
-
Beyond connectivity of locomotor circuitry-ionic and modulatory mechanisms.Prog Brain Res. 2010;187:99-110. doi: 10.1016/B978-0-444-53613-6.00007-1. Prog Brain Res. 2010. PMID: 21111203 Review.
Cited by
-
Spinal cord injury significantly alters the properties of reticulospinal neurons: delayed repolarization mediated by potassium channels.J Neurophysiol. 2023 Nov 1;130(5):1265-1281. doi: 10.1152/jn.00251.2023. Epub 2023 Oct 11. J Neurophysiol. 2023. PMID: 37820016 Free PMC article.
-
Heterologous functional expression of ascidian Nav1 channels and close relationship with the evolutionary ancestor of vertebrate Nav channels.J Biol Chem. 2021 Jan-Jun;296:100783. doi: 10.1016/j.jbc.2021.100783. Epub 2021 May 14. J Biol Chem. 2021. PMID: 34000300 Free PMC article.
References
-
- Batueva IV, Tsvetkov EA, Buchanan JT, Veselkin NP, 1996. Investigation of voltage- gated currents in the isolated neurons of river lamprey Lampetra fluviatilis spinal cord. J Evol 217–231. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

