The Paraventricular Hypothalamus Regulates Satiety and Prevents Obesity via Two Genetically Distinct Circuits
- PMID: 30879785
- PMCID: PMC6508999
- DOI: 10.1016/j.neuron.2019.02.028
The Paraventricular Hypothalamus Regulates Satiety and Prevents Obesity via Two Genetically Distinct Circuits
Abstract
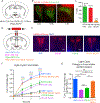
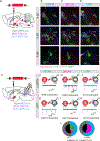
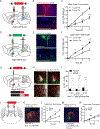
SIM1-expressing paraventricular hypothalamus (PVH) neurons are key regulators of energy balance. Within the PVHSIM1 population, melanocortin-4 receptor-expressing (PVHMC4R) neurons are known to regulate satiety and bodyweight, yet they account for only half of PVHSIM1 neuron-mediated regulation. Here we report that PVH prodynorphin-expressing (PVHPDYN) neurons, which notably lack MC4Rs, function independently and additively with PVHMC4R neurons to account for the totality of PVHSIM1 neuron-mediated satiety. Moreover, PVHPDYN neurons are necessary for prevention of obesity in an independent but equipotent manner to PVHMC4R neurons. While PVHPDYN and PVHMC4R neurons both project to the parabrachial complex (PB), they synaptically engage distinct efferent nodes, the pre-locus coeruleus (pLC), and central lateral parabrachial nucleus (cLPBN), respectively. PB-projecting PVHPDYN neurons, like PVHMC4R neurons, receive input from interoceptive ARCAgRP neurons, respond to caloric state, and are sufficient and necessary to control food intake. This expands the CNS satiety circuitry to include two non-overlapping PVH to hindbrain circuits.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors declare no competing interests.
Figures


Comment in
-
Parallel Paths in PVH Control of Feeding.Neuron. 2019 May 8;102(3):514-516. doi: 10.1016/j.neuron.2019.04.026. Neuron. 2019. PMID: 31071283
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

