Surface-Binding to Cardiolipin Nanodomains Triggers Cytochrome c Pro-apoptotic Peroxidase Activity via Localized Dynamics
- PMID: 30879887
- PMCID: PMC6615723
- DOI: 10.1016/j.str.2019.02.007
Surface-Binding to Cardiolipin Nanodomains Triggers Cytochrome c Pro-apoptotic Peroxidase Activity via Localized Dynamics
Abstract
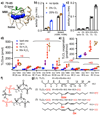
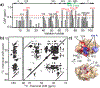
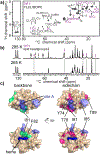
The peroxidation of cardiolipins by reactive oxygen species, which is regulated and enhanced by cytochrome c (cyt c), is a critical signaling event in mitochondrial apoptosis. We probe the molecular underpinnings of this mitochondrial death signal through structural and functional studies of horse heart cyt c binding to mixed-lipid membranes containing cardiolipin with mono- and polyunsaturated acyl chains. Lipidomics reveal the selective oxidation of polyunsaturated fatty acid (PUFA) cardiolipin (CL), while multidimensional solid-state NMR probes the structure and dynamics of the membrane and the peripherally bound protein. The hydrophilic milieu at the membrane interface stabilizes a native-like fold, but also leads to localized flexibility at the membrane-interacting protein face. PUFA CL acts as both a preferred substrate and a dynamic regulator by affecting the dynamics of the cyt c N70-I85 Ω loop, which covers the heme cavity.
Keywords: PUFA; apoptosis; cardiolipin; cytochrome c; lipidomics; membrane oxidation; membrane protein; mitochondrial protein; protein structure and dynamics; solid-state NMR.
Copyright © 2019 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Interests
The authors declare no competing interests.
Figures



Similar articles
-
Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids.J Mol Biol. 2021 Jul 23;433(15):167057. doi: 10.1016/j.jmb.2021.167057. Epub 2021 May 24. J Mol Biol. 2021. PMID: 34033821 Free PMC article.
-
Structural Changes and Proapoptotic Peroxidase Activity of Cardiolipin-Bound Mitochondrial Cytochrome c.Biophys J. 2015 Nov 3;109(9):1873-84. doi: 10.1016/j.bpj.2015.09.016. Biophys J. 2015. PMID: 26536264 Free PMC article.
-
Topography of tyrosine residues and their involvement in peroxidation of polyunsaturated cardiolipin in cytochrome c/cardiolipin peroxidase complexes.Biochim Biophys Acta. 2011 Sep;1808(9):2147-55. doi: 10.1016/j.bbamem.2011.04.009. Epub 2011 Apr 29. Biochim Biophys Acta. 2011. PMID: 21550335 Free PMC article.
-
Apoptotic interactions of cytochrome c: redox flirting with anionic phospholipids within and outside of mitochondria.Biochim Biophys Acta. 2006 May-Jun;1757(5-6):648-59. doi: 10.1016/j.bbabio.2006.03.002. Epub 2006 Mar 31. Biochim Biophys Acta. 2006. PMID: 16740248 Review.
-
The "pro-apoptotic genies" get out of mitochondria: oxidative lipidomics and redox activity of cytochrome c/cardiolipin complexes.Chem Biol Interact. 2006 Oct 27;163(1-2):15-28. doi: 10.1016/j.cbi.2006.04.019. Epub 2006 May 12. Chem Biol Interact. 2006. PMID: 16797512 Review.
Cited by
-
Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids.J Mol Biol. 2021 Jul 23;433(15):167057. doi: 10.1016/j.jmb.2021.167057. Epub 2021 May 24. J Mol Biol. 2021. PMID: 34033821 Free PMC article.
-
Alkaline State of the Domain-Swapped Dimer of Human Cytochrome c: A Conformational Switch for Apoptotic Peroxidase Activity.J Am Chem Soc. 2022 Nov 23;144(46):21184-21195. doi: 10.1021/jacs.2c08325. Epub 2022 Nov 8. J Am Chem Soc. 2022. PMID: 36346995 Free PMC article.
-
Redox Pioneer: Professor Valerian Kagan.Antioxid Redox Signal. 2022 May;36(13-15):813-823. doi: 10.1089/ars.2021.0079. Epub 2022 Apr 21. Antioxid Redox Signal. 2022. PMID: 35072541 Free PMC article.
-
Solid-state NMR protocols for unveiling dynamics and (drug) interactions of membrane-bound proteins.Protein Sci. 2025 Apr;34(4):e70102. doi: 10.1002/pro.70102. Protein Sci. 2025. PMID: 40099898 Free PMC article.
-
Redox phospholipidomics of enzymatically generated oxygenated phospholipids as specific signals of programmed cell death.Free Radic Biol Med. 2020 Feb 1;147:231-241. doi: 10.1016/j.freeradbiomed.2019.12.028. Epub 2019 Dec 25. Free Radic Biol Med. 2020. PMID: 31883467 Free PMC article. Review.
References
-
- Abe M, Niibayashi R, Koubori S, Moriyama I, and Miyoshi H (2011). Molecular mechanisms for the induction of peroxidase activity of the cytochrome c-cardiolipin complex. Biochemistry 50, 8383–8391. - PubMed
-
- Alvarez-Paggi D, Hannibal L, Castro MA, Oviedo-Rouco S, Demicheli V, Tortora V, Tomasina F, Radi R, and Murgida DH (2017). Multifunctional cytochrome c: Learning new tricks from an old dog. Chem. Rev 117, 13382–13460. - PubMed
-
- Bachar M, Brunelle P, Tieleman DP, and Rauk A (2004). Molecular Dynamics Simulation of a Polyunsaturated Lipid Bilayer Susceptible to Lipid Peroxidation. J. Phys. Chem. B 108, 7170–7179.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

