Integrative analysis identifies DNMTs against immune-infiltrating neutrophils and dendritic cells in colorectal cancer
- PMID: 30880552
- PMCID: PMC6557545
- DOI: 10.1080/15592294.2019.1588684
Integrative analysis identifies DNMTs against immune-infiltrating neutrophils and dendritic cells in colorectal cancer
Abstract
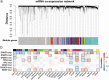
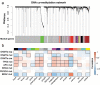
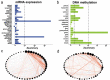
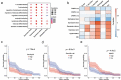
Molecular characterizations, including microsatellite instability (MSI) and the CpG island methylator phenotype (CIMP) showed strong associations in colorectal carcinoma (CRC) and provided a deeper understanding of the etiology of disease. However, the global relationship between epigenetic alternations and changes in mRNA expression in CRC remains largely undefined, especially regarding the roles of DNA methyltransferases (DNMTs). Here, we conducted a systematic network comparison to explore the global conservation between co-expressed and co-methylated modules. We successfully identified immune-related modules that were regulated by DNMTs and had strong associations with immune-infiltrating neutrophils and dendritic cells in CRC. Moreover, we found that genes in those modules were prognostic for CRC, with 97.1% (168/173) being significantly influenced by DNMTs. Thus, this study resolved an interaction between DNA methylation and mRNA expression through DNMTs. Additionally, we provided evidence that DNMTs control the global hypomethylation of oncogenes, including ALOX5AP and CSF3R that otherwise have high methylation in normal colons. Such genes were also more sensitive to DNMT changes, such as in CRC. Collectively, our analyzes provided a systems biology approach to investigate the association among different molecular phenotypes in diseases.
Keywords: Colorectal carcinoma; DNMTs; WGCNA; dendritic cell; neutrophil.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
