RecBCD, SbcCD and ExoI process a substrate created by convergent replisomes to complete DNA replication
- PMID: 30883946
- PMCID: PMC6561825
- DOI: 10.1111/mmi.14242
RecBCD, SbcCD and ExoI process a substrate created by convergent replisomes to complete DNA replication
Abstract
The accurate completion of DNA replication on the chromosome requires RecBCD and structure specific SbcCD and ExoI nucleases. However, the substrates and mechanism by which this reaction occurs remains unknown. Here we show that these completion enzymes operate on plasmid substrates containing two replisomes, but are not required for plasmids containing one replisome. Completion on the two-replisome plasmids requires RecBCD, but does not require RecA and no broken intermediates accumulate in its absence, indicating that the completion reaction occurs normally in the absence of any double-strand breaks. Further, similar to the chromosome, we show that when the normal completion reaction is prevented, an aberrant RecA-mediated recombination process leads to amplifications that drive most of the instabilities associated with the two-replisome substrates. The observations imply that the substrate SbcCD, ExoI and RecBCD act upon in vivo is created specifically by two convergent replisomes, and demonstrate that the function of RecBCD in completing replication is independent of double-strand break repair, and likely promotes joining of the strands of the convergent replication forks.
© 2019 John Wiley & Sons Ltd.
Figures

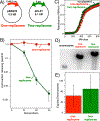
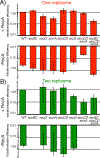
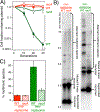

Similar articles
-
Completion of DNA replication in Escherichia coli.Proc Natl Acad Sci U S A. 2014 Nov 18;111(46):16454-9. doi: 10.1073/pnas.1415025111. Epub 2014 Nov 3. Proc Natl Acad Sci U S A. 2014. PMID: 25368150 Free PMC article.
-
SbcC-SbcD and ExoI process convergent forks to complete chromosome replication.Proc Natl Acad Sci U S A. 2018 Jan 9;115(2):349-354. doi: 10.1073/pnas.1715960114. Epub 2017 Dec 5. Proc Natl Acad Sci U S A. 2018. PMID: 29208713 Free PMC article.
-
chi sequences switch the RecBCD helicase-nuclease complex from degradative to replicative modes during the completion of DNA replication.J Biol Chem. 2023 Mar;299(3):103013. doi: 10.1016/j.jbc.2023.103013. Epub 2023 Feb 11. J Biol Chem. 2023. PMID: 36781123 Free PMC article.
-
RecBCD is required to complete chromosomal replication: Implications for double-strand break frequencies and repair mechanisms.DNA Repair (Amst). 2015 Aug;32:86-95. doi: 10.1016/j.dnarep.2015.04.018. Epub 2015 May 2. DNA Repair (Amst). 2015. PMID: 26003632 Free PMC article. Review.
-
Single-strand gap repair involves both RecF and RecBCD pathways.Curr Genet. 2016 Aug;62(3):519-21. doi: 10.1007/s00294-016-0575-5. Epub 2016 Feb 13. Curr Genet. 2016. PMID: 26874520 Review.
Cited by
-
The Roles of Bacterial DNA Double-Strand Break Repair Proteins in Chromosomal DNA Replication.FEMS Microbiol Rev. 2020 May 1;44(3):351-368. doi: 10.1093/femsre/fuaa009. FEMS Microbiol Rev. 2020. PMID: 32286623 Free PMC article. Review.
-
Recombination Mediator Proteins: Misnomers That Are Key to Understanding the Genomic Instabilities in Cancer.Genes (Basel). 2022 Feb 27;13(3):437. doi: 10.3390/genes13030437. Genes (Basel). 2022. PMID: 35327990 Free PMC article. Review.
-
Role for DNA double strand end-resection activity of RecBCD in control of aberrant chromosomal replication initiation in Escherichia coli.Nucleic Acids Res. 2022 Aug 26;50(15):8643-8657. doi: 10.1093/nar/gkac670. Nucleic Acids Res. 2022. PMID: 35929028 Free PMC article.
-
Under-Replicated DNA: The Byproduct of Large Genomes?Cancers (Basel). 2020 Sep 25;12(10):2764. doi: 10.3390/cancers12102764. Cancers (Basel). 2020. PMID: 32992928 Free PMC article. Review.
-
Bacterial Argonaute nucleases reveal different modes of DNA targeting in vitro and in vivo.Nucleic Acids Res. 2023 Jun 9;51(10):5106-5124. doi: 10.1093/nar/gkad290. Nucleic Acids Res. 2023. PMID: 37094066 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

