PTEN arginine methylation by PRMT6 suppresses PI3K-AKT signaling and modulates pre-mRNA splicing
- PMID: 30886105
- PMCID: PMC6452682
- DOI: 10.1073/pnas.1811028116
PTEN arginine methylation by PRMT6 suppresses PI3K-AKT signaling and modulates pre-mRNA splicing
Abstract
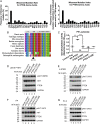
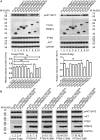
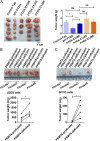
Arginine methylation is a ubiquitous posttranslational modification that regulates critical cellular processes including signal transduction and pre-mRNA splicing. Here, we report that the tumor-suppressor PTEN is methylated by protein arginine methyltransferase 6 (PRMT6). Mass-spectrometry analysis reveals that PTEN is dimethylated at arginine 159 (R159). We found that PTEN is mutated at R159 in cancers, and the PTEN mutant R159K loses its capability to inhibit the PI3K-AKT cascade. Furthermore, PRMT6 is physically associated with PTEN, promotes asymmetrical dimethylation of PTEN, and regulates the PI3K-AKT cascade through PTEN R159 methylation. In addition, using transcriptome analyses, we found that PTEN R159 methylation is involved in modulation of pre-mRNA alternative splicing. Our results demonstrate that PTEN is functionally regulated by arginine methylation. We propose that PTEN arginine methylation modulates pre-mRNA alternative splicing and influences diverse physiologic processes.
Keywords: PI3K–AKT cascade; PRMT6; PTEN; arginine methylation; pre-mRNA splicing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Cigarette smoke extract induces airway epithelial cell death via repressing PRMT6/AKT signaling.Aging (Albany NY). 2020 Dec 1;12(23):24301-24317. doi: 10.18632/aging.202210. Epub 2020 Dec 1. Aging (Albany NY). 2020. PMID: 33260152 Free PMC article.
-
Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role.Nat Commun. 2018 Jun 19;9(1):2392. doi: 10.1038/s41467-018-04760-1. Nat Commun. 2018. PMID: 29921876 Free PMC article.
-
Long Noncoding RNA lncARSR Promotes Doxorubicin Resistance in Hepatocellular Carcinoma via Modulating PTEN-PI3K/Akt Pathway.J Cell Biochem. 2017 Dec;118(12):4498-4507. doi: 10.1002/jcb.26107. Epub 2017 Jun 9. J Cell Biochem. 2017. PMID: 28464252
-
Gene alterations in the PI3K/PTEN/AKT pathway as a mechanism of drug-resistance (review).Int J Oncol. 2012 Mar;40(3):639-44. doi: 10.3892/ijo.2011.1312. Epub 2011 Dec 20. Int J Oncol. 2012. PMID: 22200790 Review.
-
Targeting the translational apparatus to improve leukemia therapy: roles of the PI3K/PTEN/Akt/mTOR pathway.Leukemia. 2011 Jul;25(7):1064-79. doi: 10.1038/leu.2011.46. Epub 2011 Mar 25. Leukemia. 2011. PMID: 21436840 Review.
Cited by
-
Exploiting epigenetic dependencies in ovarian cancer therapy.Int J Cancer. 2021 Nov 15;149(10):1732-1743. doi: 10.1002/ijc.33727. Epub 2021 Aug 6. Int J Cancer. 2021. PMID: 34213777 Free PMC article. Review.
-
The Expression of Proto-Oncogene ETS-Related Gene (ERG) Plays a Central Role in the Oncogenic Mechanism Involved in the Development and Progression of Prostate Cancer.Int J Mol Sci. 2022 Apr 26;23(9):4772. doi: 10.3390/ijms23094772. Int J Mol Sci. 2022. PMID: 35563163 Free PMC article. Review.
-
Arginine methylation in the epithelial-to-mesenchymal transition.FEBS J. 2022 Dec;289(23):7292-7303. doi: 10.1111/febs.16152. Epub 2021 Aug 15. FEBS J. 2022. PMID: 34358413 Free PMC article. Review.
-
PRMT1 and PRMT5: on the road of homologous recombination and non-homologous end joining.Genome Instab Dis. 2023 Aug;4(4):197-209. doi: 10.1007/s42764-022-00095-w. Epub 2022 Dec 7. Genome Instab Dis. 2023. PMID: 37663901 Free PMC article.
-
PTEN Methylation by NSD2 Controls Cellular Sensitivity to DNA Damage.Cancer Discov. 2019 Sep;9(9):1306-1323. doi: 10.1158/2159-8290.CD-18-0083. Epub 2019 Jun 19. Cancer Discov. 2019. PMID: 31217297 Free PMC article.
References
-
- Li J, et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science. 1997;275:1943–1947. - PubMed
-
- Steck PA, et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat Genet. 1997;15:356–362. - PubMed
-
- Liaw D, et al. Germline mutations of the PTEN gene in Cowden disease, an inherited breast and thyroid cancer syndrome. Nat Genet. 1997;16:64–67. - PubMed
-
- Hobert JA, Eng C. PTEN hamartoma tumor syndrome: An overview. Genet Med. 2009;11:687–694. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

