Simulations of Protein Adsorption on Nanostructured Surfaces
- PMID: 30886353
- PMCID: PMC6423022
- DOI: 10.1038/s41598-019-40920-z
Simulations of Protein Adsorption on Nanostructured Surfaces
Abstract
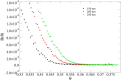
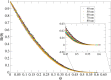
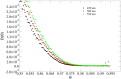
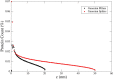
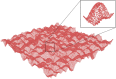
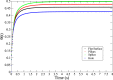
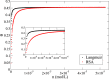
Recent technological advances have allowed the development of a new generation of nanostructured materials, such as those displaying both mechano-bactericidal activity and substrata that favor the growth of mammalian cells. Nanomaterials that come into contact with biological media such as blood first interact with proteins, hence understanding the process of adsorption of proteins onto these surfaces is highly important. The Random Sequential Adsorption (RSA) model for protein adsorption on flat surfaces was modified to account for nanostructured surfaces. Phenomena related to the nanofeature geometry have been revealed during the modelling process; e.g., convex geometries can lead to lower steric hindrance between particles, and hence higher degrees of surface coverage per unit area. These properties become more pronounced when a decrease in the size mismatch between the proteins and the surface nanostructures occurs. This model has been used to analyse the adsorption of human serum albumin (HSA) on a nano-structured black silicon (bSi) surface. This allowed the Blocking Function (the rate of adsorption) to be evaluated. The probability of the protein to adsorb as a function of the occupancy was also calculated.
Conflict of interest statement
The authors declare no competing interests.
Figures








References
-
- Rahman, F. Nanostructures in Electronics and Photonics, 1 edition edn. (Pan Stanford, Singapore, 2008).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

