Curcumin Analogue C1 Promotes Hex and Gal Recruitment to the Plasma Membrane via mTORC1-Independent TFEB Activation
- PMID: 30889901
- PMCID: PMC6471159
- DOI: 10.3390/ijms20061363
Curcumin Analogue C1 Promotes Hex and Gal Recruitment to the Plasma Membrane via mTORC1-Independent TFEB Activation
Abstract
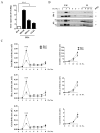
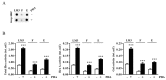
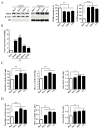
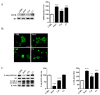
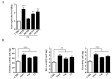
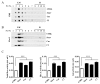
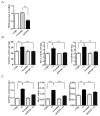
The monocarbonyl analogue of curcumin (1E,4E)-1,5-Bis(2-methoxyphenyl)penta-1,4-dien-3-one (C1) has been used as a specific activator of the master gene transcription factor EB (TFEB) to correlate the activation of this nuclear factor with the increased activity of lysosomal glycohydrolases and their recruitment to the cell surface. The presence of active lysosomal glycohydrolases associated with the lipid microdomains has been extensively demonstrated, and their role in glycosphingolipid (GSL) remodeling in both physiological and pathological conditions, such as neurodegenerative disorders, has been suggested. Here, we demonstrate that Jurkat cell stimulation elicits TFEB nuclear translocation and an increase of both the expression of hexosaminidase subunit beta (HEXB), hexosaminidase subunit alpha (HEXA), and galactosidase beta 1 (GLB1) genes, and the recruitment of β-hexosaminidase (Hex, EC 3.2.1.52) and β-galactosidase (Gal, EC 3.2.1.23) on lipid microdomains. Treatment of Jurkat cells with the curcumin analogue C1 also resulted in an increase of both lysosomal glycohydrolase activity and their targeting to the cell surface. Similar effects of C1 on lysosomal glycohydrolase expression and their recruitment to lipid microdomains was observed by treating the SH-SY5Y neuroblastoma cell line; the effects of C1 treatment were abolished by TFEB silencing. Together, these results clearly demonstrate the existence of a direct link between TFEB nuclear translocation and the transport of Hex and Gal from lysosomes to the plasma membrane.
Keywords: TFEB; curcumin; curcumin analogue C1; lysosomal glycohydrolases; plasma membrane-associated glycohydrolases.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
TFEB activation promotes the recruitment of lysosomal glycohydrolases β-hexosaminidase and β-galactosidase to the plasma membrane.Biochem Biophys Res Commun. 2013 Oct 18;440(2):251-7. doi: 10.1016/j.bbrc.2013.09.060. Epub 2013 Sep 20. Biochem Biophys Res Commun. 2013. PMID: 24055709
-
The FACT complex facilitates expression of lysosomal and antioxidant genes through binding to TFEB and TFE3.Autophagy. 2022 Oct;18(10):2333-2349. doi: 10.1080/15548627.2022.2029671. Epub 2022 Mar 1. Autophagy. 2022. PMID: 35230915 Free PMC article.
-
Glycohydrolases β-hexosaminidase and β-galactosidase are associated with lipid microdomains of Jurkat T-lymphocytes.Biochimie. 2012 Mar;94(3):684-94. doi: 10.1016/j.biochi.2011.09.021. Epub 2011 Sep 29. Biochimie. 2012. PMID: 21978926
-
TFEB at a glance.J Cell Sci. 2016 Jul 1;129(13):2475-81. doi: 10.1242/jcs.146365. Epub 2016 Jun 1. J Cell Sci. 2016. PMID: 27252382 Free PMC article. Review.
-
Lipophagy and Lipolysis Status in Lipid Storage and Lipid Metabolism Diseases.Int J Mol Sci. 2020 Aug 25;21(17):6113. doi: 10.3390/ijms21176113. Int J Mol Sci. 2020. PMID: 32854299 Free PMC article. Review.
Cited by
-
Bioactivity Evaluation of a Novel Formulated Curcumin.Nutrients. 2019 Dec 6;11(12):2982. doi: 10.3390/nu11122982. Nutrients. 2019. PMID: 31817577 Free PMC article.
-
Targeting GM2 Ganglioside Accumulation in Dementia: Current Therapeutic Approaches and Future Directions.Curr Mol Med. 2024;24(11):1329-1345. doi: 10.2174/0115665240264547231017110613. Curr Mol Med. 2024. PMID: 37877564 Review.
-
Transcription factor EB agonists from natural products for treating human diseases with impaired autophagy-lysosome pathway.Chin Med. 2020 Nov 23;15(1):123. doi: 10.1186/s13020-020-00402-1. Chin Med. 2020. PMID: 33292395 Free PMC article. Review.
-
SNX8 enables lysosome reformation and reverses lysosomal storage disorder.Nat Commun. 2024 Mar 22;15(1):2553. doi: 10.1038/s41467-024-46705-x. Nat Commun. 2024. PMID: 38519472 Free PMC article.
-
Serum Cytokine Profile, Beta-Hexosaminidase A Enzymatic Activity and GM2 Ganglioside Levels in the Plasma of a Tay-Sachs Disease Patient after Cord Blood Cell Transplantation and Curcumin Administration: A Case Report.Life (Basel). 2021 Sep 24;11(10):1007. doi: 10.3390/life11101007. Life (Basel). 2021. PMID: 34685379 Free PMC article.
References
-
- Encarnação M., Espada L., Escrevente C., Mateus D., Ramalho J., Michelet X., Santarino I., Hsu V.W., Brenner M.B., Barral D.C., et al. A Rab3a-dependent complex essential for lysosome positioning and plasma membrane repair. J. Cell Biol. 2016;213:631–640. doi: 10.1083/jcb.201511093. - DOI - PMC - PubMed
-
- Giatromanolaki A., Kalamida D., Sivridis E., Karagounis I.V., Gatter K.C., Harris A.L., Koukourakis M.I. Increased expression of transcription factor EB (TFEB) is associated with autophagy, migratory phenotype and poor prognosis in non-small cell lung cancer. Lung Cancer. 2015;90:98–105. doi: 10.1016/j.lungcan.2015.07.008. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

